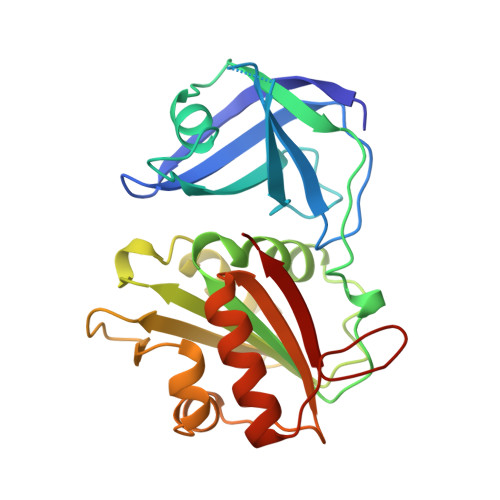
The three-dimensional structure of flavodoxin reductase from Escherichia coli at 1.7 A resolution.
Ingelman, M., Bianchi, V., Eklund, H.(1997) J Mol Biology 268: 147-157
- PubMed: 9149148
- DOI: https://doi.org/10.1006/jmbi.1997.0957
- Primary Citation Related Structures:
1FDR - PubMed Abstract:
Flavodoxin reductase from Escherichia coli is an FAD-containing oxidoreductase that transports electrons between flavodoxin or ferredoxin and NADPH. Together with flavodoxin, the enzyme is involved in the reductive activation of three E. coli enzymes: cobalamin-dependent methionine synthase, pyruvate formate lyase and anaerobic ribonucleotide reductase. An additional function for the oxidoreductase appears to be to protect the bacteria against oxygen radicals. The three-dimensional structure of flavodoxin reductase has been solved by multiple isomorphous replacement, and has been refined at 1.7 A to an R-value of 18.4% and Rfree 24.8%. The monomeric molecule contains one beta-sandwich FAD domain and an alpha/beta NADP domain. The overall structure is similar to other reductases of the NADP-ferredoxin reductase family in spite of the low sequence similarities within the family. Flavodoxin reductase lacks the loop which is involved in the binding of the adenosine moiety of FAD in other FAD binding enzymes of the superfamily but is missing in the FMN binding phthalate dioxygenase reductase. Instead of this loop, the adenine interacts with an extra tryptophan at the C terminus. The FAD in flavodoxin reductase has an unusual bent conformation with a hydrogen bond between the adenine and the isoalloxazine. This is probably the cause of the unusual spectrum of the enzyme. There is a pronounced cleft close to the isoalloxazine that appears to be well suited for binding of flavodoxin/ferredoxin. Two extra short strands of the NADP-binding domain probably act as an anchor point for the binding of flavodoxin.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala Biomedical Center.
Organizational Affiliation: