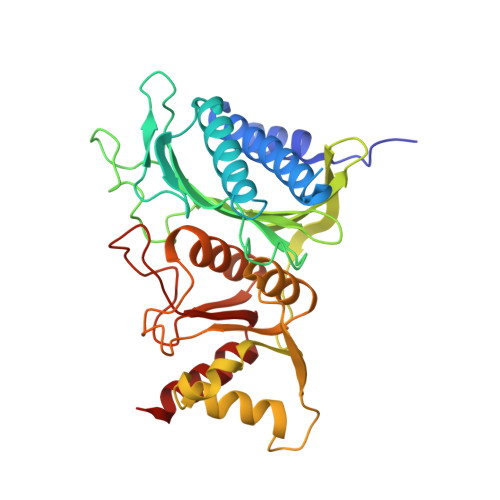
Crystal structure of fructose-1,6-bisphosphatase complexed with fructose 6-phosphate, AMP, and magnesium.
Ke, H.M., Zhang, Y.P., Lipscomb, W.N.(1990) Proc Natl Acad Sci U S A 87: 5243-5247
- PubMed: 2164670 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.87.14.5243
- Primary Citation Related Structures:
1FBP - PubMed Abstract:
The crystal structure of fructose-1,6-bisphosphatase (EC 3.1.3.11) complexed with fructose 6-phosphate, AMP, and Mg2+ has been solved by the molecular replacement method and refined at 2.5-A resolution to a R factor of 0.215, with root-mean-square deviations of 0.013 A and 3.5 degrees for bond lengths and bond angles, respectively. No solvent molecules have been included in the refinement. This structure shows large quaternary and tertiary conformational changes from the structures of the unligated enzyme or its fructose 2,6-bisphosphate complex, but the secondary structures remain essentially the same. Dimer C3-C4 of the enzyme-fructose 6-phosphate-AMP-Mg2+ complex twists about 19 degrees relative to the same dimer of the enzyme-fructose 2,6-bisphosphate complex if their C1-C2 dimers are superimposed on one another. Nevertheless, many interfacial interactions between dimers of C1-C2 and C3-C4 are conserved after quaternary structure changes occur. Residues of the AMP domain (residues 6-200) show large migrations of C alpha atoms relative to barely significant positional changes of the FBP domain (residues 201-335).
- Gibbs Chemical Laboratory, Harvard University, Cambridge, MA 02138.
Organizational Affiliation: