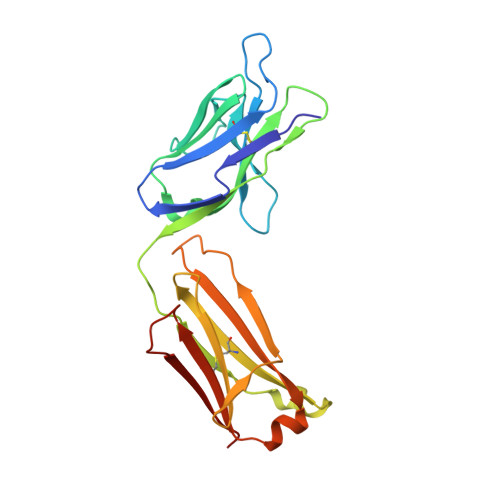
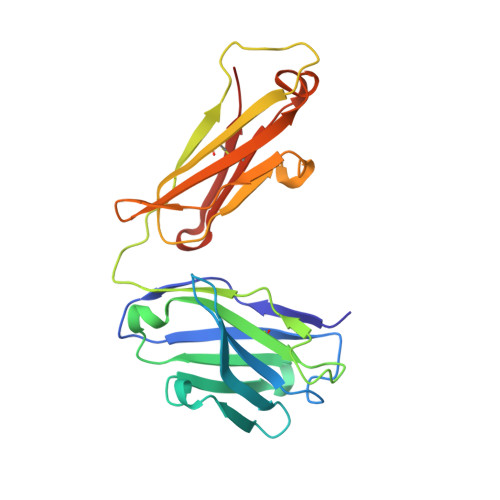
Structure of the Fab fragment from F124, a monoclonal antibody specific for hepatitis B surface antigen.
Saul, F.A., Vulliez-Le Normand, B., Passafiume, M., Riottot, M.M., Bentley, G.A.(2000) Acta Crystallogr D Biol Crystallogr 56: 945-951
- PubMed: 10944330 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444900008088
- Primary Citation Related Structures:
1F11 - PubMed Abstract:
The crystal structure of the Fab fragment from the monoclonal anti-preS2 antibody F124 (IgG1,kappa) has been solved by molecular replacement and refined at 3.0 A resolution. The Fab crystallizes with two independent molecules in the asymmetric unit. F124 recognizes an epitope contained within the preS2 segment between residues 120 and 132 of the surface antigen of hepatitis B virus. The antibody shows a high affinity for the glycan N-linked to Asn123, but it also cross-reacts with the non-glycosylated peptide fragment 120-132. Although crystallization was performed in the presence of an eightfold excess of the cross-reactive peptide, no evidence for the ligand was found in the antigen-binding site, which is close to a neighbouring molecule in the crystal lattice. The antigen-binding site has a groove-like topology which is modulated with pocket-like cavities. It is characterized by a large number of tyrosine and aspartate residues. The importance of germ-line mutations at the binding site is discussed.
- Unité d'Immunologie Structurale, CNRS URA 2185, Institut Pasteur, 25 Rue du Dr Roux, 75724 Paris, France.
Organizational Affiliation: