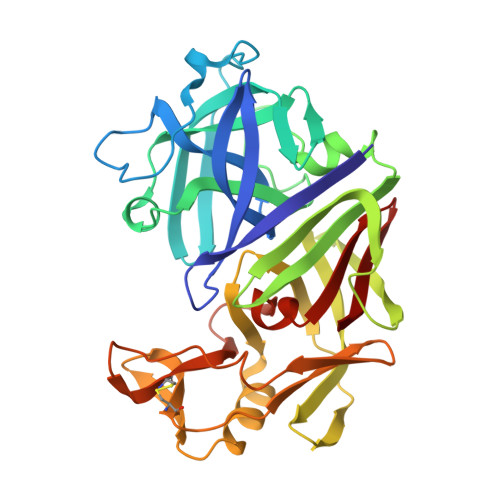
Direct observation by X-ray analysis of the tetrahedral intermediate of aspartic proteinases.
Veerapandian, B., Cooper, J.B., Sali, A., Blundell, T.L., Rosati, R.L., Dominy, B.W., Damon, D.B., Hoover, D.J.(1992) Protein Sci 1: 322-328
- PubMed: 1304340 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560010303
- Primary Citation Related Structures:
1EPO - PubMed Abstract:
We report the X-ray analysis at 2.0 A resolution for crystals of the aspartic proteinase endothiapepsin (EC 3.4.23.6) complexed with a potent difluorostatone-containing tripeptide renin inhibitor (CP-81,282). The scissile bond surrogate, an electrophilic ketone, is hydrated in the complex. The pro-(R) (statine-like) hydroxyl of the tetrahedral carbonyl hydrate is hydrogen-bonded to both active-site aspartates 32 and 215 in the position occupied by a water in the native enzyme. The second hydroxyl oxygen of the hydrate is hydrogen-bonded only to the outer oxygen of Asp 32. These experimental data provide a basis for a model of the tetrahedral intermediate in aspartic proteinase-mediated cleavage of the amide bond. This indicates a mechanism in which Asp 32 is the proton donor and Asp 215 carboxylate polarizes a bound water for nucleophilic attack. The mechanism involves a carboxylate (Asp 32) that is stabilized by extensive hydrogen bonding, rather than an oxyanion derivative of the peptide as in serine proteinase catalysis.
- Department of Crystallography, Birkbeck College, London, UK.
Organizational Affiliation: