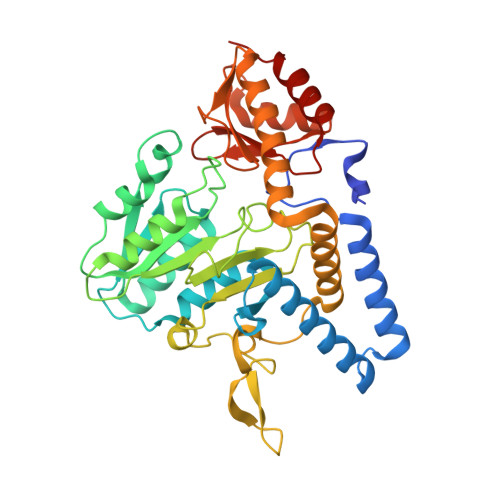
Crystal structure of the cystine C-S lyase from Synechocystis: stabilization of cysteine persulfide for FeS cluster biosynthesis.
Clausen, T., Kaiser, J.T., Steegborn, C., Huber, R., Kessler, D.(2000) Proc Natl Acad Sci U S A 97: 3856-3861
- PubMed: 10760256
- DOI: https://doi.org/10.1073/pnas.97.8.3856
- Primary Citation Related Structures:
1ELQ, 1ELU - PubMed Abstract:
FeS clusters are versatile cofactors of a variety of proteins, but the mechanisms of their biosynthesis are still unknown. The cystine C-S lyase from Synechocystis has been identified as a participant in ferredoxin FeS cluster formation. Herein, we report on the crystal structure of the lyase and of a complex with the reaction products of cystine cleavage at 1.8- and 1.55-A resolution, respectively. The sulfur-containing product was unequivocally identified as cysteine persulfide. The reactive persulfide group is fixed by a hydrogen bond to His-114 in the center of a hydrophobic pocket and is thereby shielded from the solvent. Binding and stabilization of the cysteine persulfide represent an alternative to the generation of a protein-bound persulfide by NifS-like proteins and point to the general importance of persulfidic compounds for FeS cluster assembly.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried, Germany. clausen@biochem.mpg.de
Organizational Affiliation: