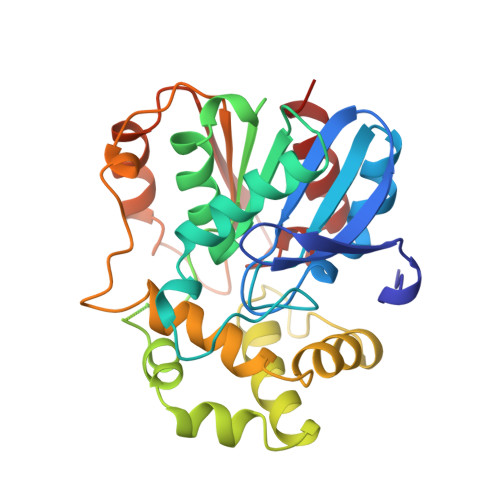
The x-ray structure of epoxide hydrolase from Agrobacterium radiobacter AD1. An enzyme to detoxify harmful epoxides.
Nardini, M., Ridder, I.S., Rozeboom, H.J., Kalk, K.H., Rink, R., Janssen, D.B., Dijkstra, B.W.(1999) J Biological Chem 274: 14579-14586
- PubMed: 10329649 Search on PubMed
- Primary Citation Related Structures:
1EHY - PubMed Abstract:
Epoxide hydrolases catalyze the cofactor-independent hydrolysis of reactive and toxic epoxides. They play an essential role in the detoxification of various xenobiotics in higher organisms and in the bacterial degradation of several environmental pollutants. The first x-ray structure of one of these, from Agrobacterium radiobacter AD1, has been determined by isomorphous replacement at 2.1-A resolution. The enzyme shows a two-domain structure with the core having the alpha/beta hydrolase-fold topology. The catalytic residues, Asp107 and His275, are located in a predominantly hydrophobic environment between the two domains. A tunnel connects the back of the active-site cavity with the surface of the enzyme and provides access to the active site for the catalytic water molecule, which in the crystal structure, has been found at hydrogen bond distance to His275. Because of a crystallographic contact, the active site has become accessible for the Gln134 side chain, which occupies a position mimicking a bound substrate. The structure suggests Tyr152/Tyr215 as the residues involved in substrate binding, stabilization of the transition state, and possibly protonation of the epoxide oxygen.
- Laboratory of Biophysical Chemistry and BIOSON Research Institute, Department of Chemistry, University of Groningen, Nijenborgh 4, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: