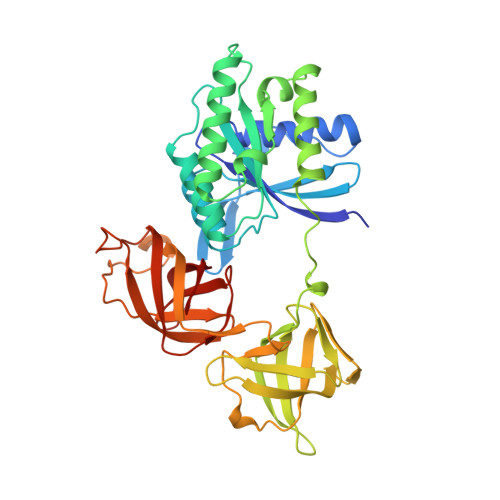
Crystal structure of intact elongation factor EF-Tu from Escherichia coli in GDP conformation at 2.05 A resolution.
Song, H., Parsons, M.R., Rowsell, S., Leonard, G., Phillips, S.E.(1999) J Mol Biology 285: 1245-1256
- PubMed: 9918724 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2387
- Primary Citation Related Structures:
1EFC - PubMed Abstract:
The crystal structure of intact elongation factor Tu (EF-Tu) from Escherichia coli in GDP-bound conformation has been determined using a combination of multiple isomorphous replacement (MIR) and multiwavelength anomalous diffraction (MAD) methods. The current atomic model has been refined to a crystallographic R factor of 20.3 % and free R-factor of 26.8 % in the resolution range of 10-2.05 A. The protein consists of three domains: domain 1 has an alpha/beta structure; while domain 2 and domain 3 are beta-barrel structures. Although the global fold of the current model is similar to those of published structures, the secondary structural assignment has been improved due to the high quality of the current model. The switch I region (residues 40-62) is well ordered in this structure. Comparison with the structure of EF-Tu in GDP-bound form from Thermus aquaticus shows that although the individual domain structures are similar in these two structures, the orientation of domains changes significantly. Interactions between domains 1 and 3 in our E. coli EF-Tu-GDP complex are quite different from those of EF-Tu with bound GTP from T. aquaticus, due to the domain rearrangement upon GTP binding. The binding sites of the Mg2+ and guanine nucleotide are revealed in detail. Two water molecules that co-ordinate the Mg2+ have been identified to be well conserved in the GDP and GTP-bound forms of EF-Tu structures, as well as in the structure of Ras p21 with bound GDP. Comparisons of the Mg2+ binding site with other guanine nucleotide binding proteins in GDP-bound forms show that the Mg2+ co-ordination patterns are well preserved among these structures.
- School of Biochemistry & Molecular Biology, University of Leeds, Leeds, LS2 9JT, UK.
Organizational Affiliation: