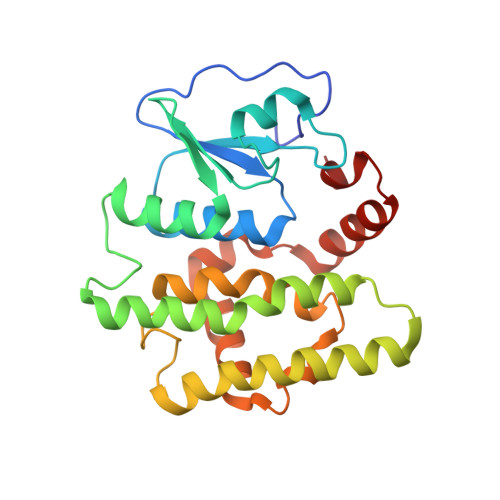
Identification, characterization, and crystal structure of the Omega class glutathione transferases.
Board, P.G., Coggan, M., Chelvanayagam, G., Easteal, S., Jermiin, L.S., Schulte, G.K., Danley, D.E., Hoth, L.R., Griffor, M.C., Kamath, A.V., Rosner, M.H., Chrunyk, B.A., Perregaux, D.E., Gabel, C.A., Geoghegan, K.F., Pandit, J.(2000) J Biological Chem 275: 24798-24806
- PubMed: 10783391 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M001706200
- Primary Citation Related Structures:
1EEM - PubMed Abstract:
A new class of glutathione transferases has been discovered by analysis of the expressed sequence tag data base and sequence alignment. Glutathione S-transferases (GSTs) of the new class, named Omega, exist in several mammalian species and Caenorhabditis elegans. In humans, GSTO 1-1 is expressed in most tissues and exhibits glutathione-dependent thiol transferase and dehydroascorbate reductase activities characteristic of the glutaredoxins. The structure of GSTO 1-1 has been determined at 2.0-A resolution and has a characteristic GST fold (Protein Data Bank entry code ). The Omega class GSTs exhibit an unusual N-terminal extension that abuts the C terminus to form a novel structural unit. Unlike other mammalian GSTs, GSTO 1-1 appears to have an active site cysteine that can form a disulfide bond with glutathione.
- Molecular Genetics Group and Human Genetics Group, John Curtin School of Medical Research, Australian National University, Canberra, Australian Capital Territory 2601, Australia.
Organizational Affiliation: