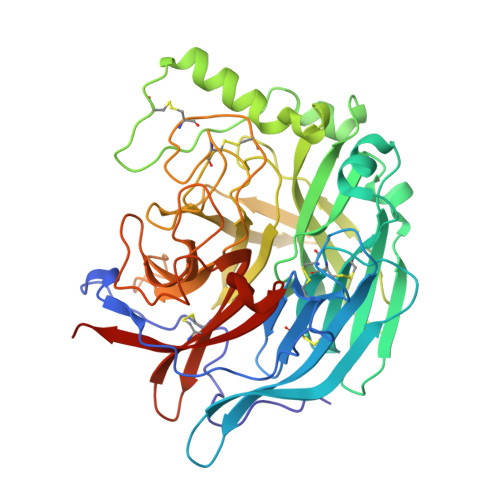
Crystal Structure of the Multifunctional Paramyxovirus Hemagglutinin-Neuraminidase
Crennell, S., Takimoto, T., Portner, A., Taylor, G.(2000) Nat Struct Biol 7: 1068
- PubMed: 11062565 Search on PubMed
- DOI: https://doi.org/10.1038/81002
- Primary Citation Related Structures:
1E8T, 1E8U, 1E8V - PubMed Abstract:
Paramyxoviruses are the main cause of respiratory disease in children. One of two viral surface glycoproteins, the hemagglutinin-neuraminidase (HN), has several functions in addition to being the major surface antigen that induces neutralizing antibodies. Here we present the crystal structures of Newcastle disease virus HN alone and in complex with either an inhibitor or with the beta-anomer of sialic acid. The inhibitor complex reveals a typical neuraminidase active site within a beta-propeller fold. Comparison of the structures of the two complexes reveal differences in the active site, suggesting that the catalytic site is activated by a conformational switch. This site may provide both sialic acid binding and hydrolysis functions since there is no evidence for a second sialic acid binding site in HN. Evidence for a single site with dual functions is examined and supported by mutagenesis studies. The structure provides the basis for the structure-based design of inhibitors for a range of paramyxovirus-induced diseases.
- Department of Biology and Biochemistry, University of Bath, Bath BA2 7AY, UK.
Organizational Affiliation: