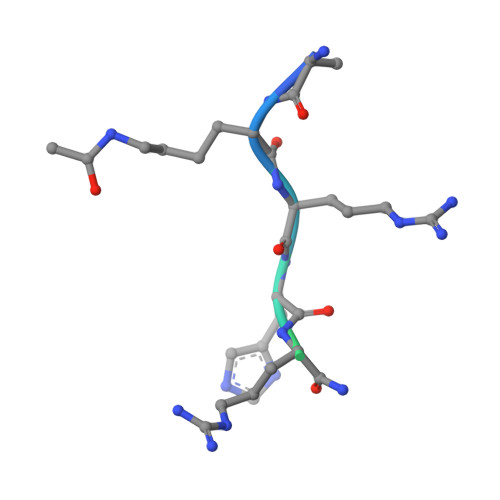
The Structural Basis for the Recognition of Acetylated Histone H4 by the Bromodomain of Histone Acetyltransferase Gcn5P
Owen, D.J., Ornaghi, P., Yang, J.C., Lowe, N., Evans, P.R., Ballario, P., Neuhaus, D., Filetici, P., Travers, A.A.(2000) EMBO J 19: 6141
- PubMed: 11080160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/19.22.6141
- Primary Citation Related Structures:
1E6I - PubMed Abstract:
The bromodomain is an approximately 110 amino acid module found in histone acetyltransferases and the ATPase component of certain nucleosome remodelling complexes. We report the crystal structure at 1.9 A resolution of the Saccharomyces cerevisiae Gcn5p bromodomain complexed with a peptide corresponding to residues 15-29 of histone H4 acetylated at the zeta-N of lysine 16. We show that this bromodomain preferentially binds to peptides containing an N:-acetyl lysine residue. Only residues 16-19 of the acetylated peptide interact with the bromodomain. The primary interaction is the N:-acetyl lysine binding in a cleft with the specificity provided by the interaction of the amide nitrogen of a conserved asparagine with the oxygen of the acetyl carbonyl group. A network of water-mediated H-bonds with protein main chain carbonyl groups at the base of the cleft contributes to the binding. Additional side chain binding occurs on a shallow depression that is hydrophobic at one end and can accommodate charge interactions at the other. These findings suggest that the Gcn5p bromodomain may discriminate between different acetylated lysine residues depending on the context in which they are displayed.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK and Centro di studio per gli Acidi Nucleici, CNR, c/o Dipartimento di Genetica e Biologia Molecolare, Università 'La Sapienza', P.le A.Moro 5, 00185 Roma, Italy.
Organizational Affiliation: