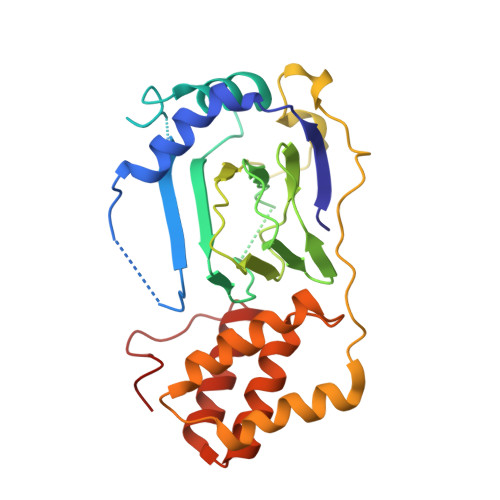
Structure of proline 3-hydroxylase. Evolution of the family of 2-oxoglutarate dependent oxygenases.
Clifton, I.J., Hsueh, L.C., Baldwin, J.E., Harlos, K., Schofield, C.J.(2001) Eur J Biochem 268: 6625-6636
- PubMed: 11737217 Search on PubMed
- DOI: https://doi.org/10.1046/j.0014-2956.2001.02617.x
- Primary Citation Related Structures:
1E5R, 1E5S - PubMed Abstract:
Iron (II)/2-oxoglutarate (2-OG)-dependent oxygenases catalyse oxidative reactions in a range of metabolic processes including the hydroxylation of proline and lysine residues during the post-translational modification of collagen. 2-OG oxygenases commonly require ascorbate for full activity. In the vitamin C deficient disease, scurvy, reduced activity of 2-OG oxygenases results in impaired formation of collagen. Here we report the crystal structure of bacterial proline 3-hydroxylase from Streptomyces sp., an enzyme which hydroxylates proline at position 3, the first of a 2-OG oxygenase catalysing oxidation of a free alpha-amino acid. Structures were obtained for the enzyme in the absence of iron (to 2.3A resolution, R=20.2%, Rfree=25.3%) and that complexed to iron (II) (to 2.4A resolution, R=19.8%, Rfree=22.6%). The structure contains conserved motifs present in other 2-OG oxygenases including a 'jelly roll' beta strand core and residues binding iron and 2-oxoglutarate, consistent with divergent evolution within the extended family. The structure differs significantly from many other 2-OG oxygenases in possessing a discrete C-terminal helical domain. Analysis of the structure suggests a model for proline binding and a mechanism for uncoupling of proline and 2-OG turnover.
- The Dyson Perrins Laboratory and the Oxford Centre for Molecular Sciences, Oxford, UK.
Organizational Affiliation: