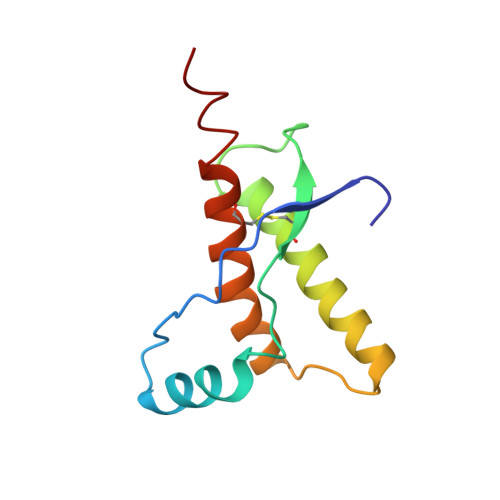
NMR Structures of Three Single-Residue Variants of the Human Prion Protein
Calzolai, L., Lysek, D.A., Guntert, P., Von Schroetter, C., Zahn, R., Riek, R., Wuthrich, K.(2000) Proc Natl Acad Sci U S A 97: 8340
- PubMed: 10900000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.97.15.8340
- Primary Citation Related Structures:
1E1G, 1E1J, 1E1P, 1E1S, 1E1U, 1E1W - PubMed Abstract:
The NMR structures of three single-amino acid variants of the C-terminal domain of the human prion protein, hPrP(121-230), are presented. In hPrP(M166V) and hPrP(R220K) the substitution is with the corresponding residue in murine PrP, and in hPrP(S170N) it is with the corresponding Syrian hamster residue. All three substitutions are in the surface region of the structure of the cellular form of PrP (PrP(C)) that is formed by the C-terminal part of helix 3, with residues 218-230, and a loop of residues 166-172. This molecular region shows high species variability and has been implicated in specific interactions with a so far not further characterized "protein X," and it is related to the species barrier for transmission of prion diseases. As expected, the three variant hPrP(121-230) structures have the same global architecture as the previously determined wild-type bovine, human, murine, and Syrian hamster prion proteins, but with the present study two localized "conformational markers" could be related with single amino acid exchanges. These are the length and quality of definition of helix 3, and the NMR-observability of the residues in the loop 166-172. Poor definition of the C-terminal part of helix 3 is characteristic for murine PrP and has now been observed also for hPrP(R220K), and NMR observation of the complete loop 166-172 has so far been unique for Syrian hamster PrP and is now also documented for hPrP(S170N).
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule, Hönggerberg, CH-8093 Zürich, Switzerland.
Organizational Affiliation: