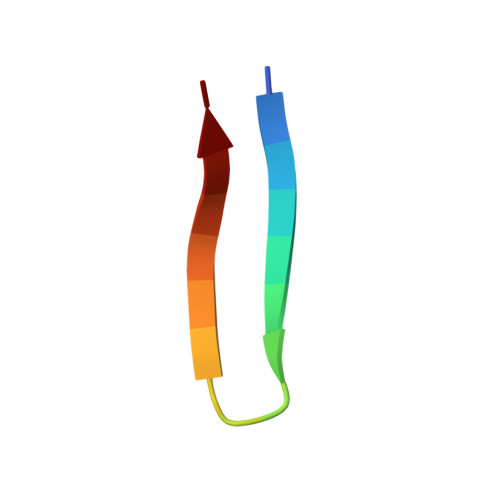
Structural Characterization of a Mutant Peptide Derived from Ubiquitin: Implications for Protein Folding.
Zerella, R., Chen, P.Y., Evans, P.A., Raine, A., Williams, D.H.(2000) Protein Sci 9: 2142
- PubMed: 11152124
- DOI: https://doi.org/10.1110/ps.9.11.2142
- Primary Citation Related Structures:
1E0Q - PubMed Abstract:
The formation of the N-terminal beta-hairpin of ubiquitin is thought to be an early event in the folding of this small protein. Previously, we have shown that a peptide corresponding to residues 1-17 of ubiquitin folds autonomously and is likely to have a native-like hairpin register. To investigate the causes of the stability of this fold, we have made mutations in the amino acids at the apex of the turn. We find that in a peptide where Thr9 is replaced by Asp, U(1-17)T9D, the native conformation is stabilized with respect to the wild-type sequence, so much so that we are able to characterize the structure of the mutant peptide fully by NMR spectroscopy. The data indicate that U(1-17)T9D peptide does indeed form a hairpin with a native-like register and a type I turn with a G1 beta-bulge, as in the full-length protein. The reason for the greater stability of the U(1-17)T9D mutant remains uncertain, but there are nuclear Overhauser effects between the side chains of Asp9 and Lys 11, which may indicate that a charge-charge interaction between these residues is responsible.
- Cambridge Centre for Molecular Recognition and University Chemical Laboratory, United Kingdom.
Organizational Affiliation: