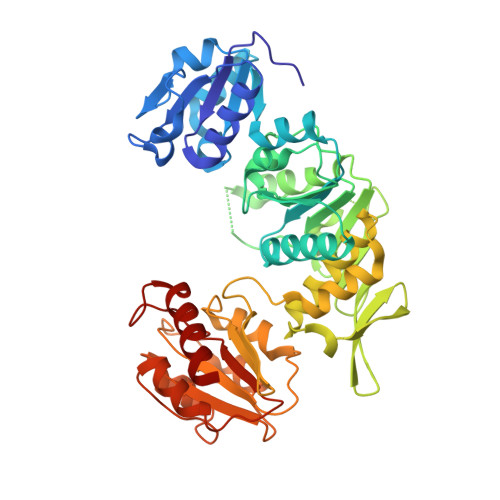
"Open" Structures of Murd: Domain Movements and Structural Similarities with Folylpolyglutamate Synthetase.
Bertrand, J., Fanchon, E., Martin, L., Chantalat, L., Auger, G., Blanot, D., Van Heijenoort, J., Dideberg, O.(2000) J Mol Biology 301: 1257
- PubMed: 10966819 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3994
- Primary Citation Related Structures:
1E0D, 1EEH - PubMed Abstract:
UDP-N-acetylmuramoyl-l-alanine:d-glutamate (MurD) ligase catalyses the addition of d-glutamate to the nucleotide precursor UDP-N-acetylmuramoyl-l-alanine (UMA). The crystal structures of Escherichia coli in the substrate-free form and MurD complexed with UMA have been determined at 2.4 A and 1.88 A resolution, respectively. The MurD structure comprises three domains each of a topology reminiscent of nucleotide-binding folds. In the two structures the C-terminal domain undergoes a large rigid-body rotation away from the N-terminal and central domains. These two "open" structures were compared with the four published "closed" structures of MurD. In addition the comparison reveals which regions are affected by the binding of UMA, ATP and d-Glu. Also we compare and discuss two structurally characterized enzymes which belong to the same ligase superfamily: MurD and folylpolyglutamate synthetase (FGS). The analysis allows the identification of key residues involved in the reaction mechanism of FGS. The determination of the two "open" conformation structures represents a new step towards the complete elucidation of the enzymatic mechanism of the MurD ligase.
- Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale Jean-Pierre Ebel (CNRS-CEA), 41, rue Jules Horowitz, Grenoble, Cedex 1, F-38027, France.
Organizational Affiliation: