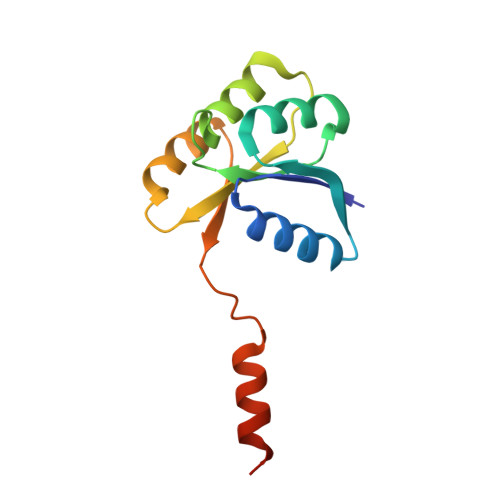
Domain swapping in the sporulation response regulator Spo0A.
Lewis, R.J., Muchova, K., Brannigan, J.A., Barak, I., Leonard, G., Wilkinson, A.J.(2000) J Mol Biology 297: 757-770
- PubMed: 10731426 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3598
- Primary Citation Related Structures:
1DZ3 - PubMed Abstract:
Adaptive responses of micro-organisms, such as chemotaxis and sporulation, are governed by two-component systems consisting of sensor kinases, that interpret environmental signals, and response regulators which activate the appropriate physiological responses. Signal transduction via response regulator proteins is mediated through transient phosphorylation of aspartic acid residues. In Spo0A, the key regulator of development (sporulation) in Bacillus, phosphorylation of the N-terminal receiver domain (N-Spo0A) at aspartate-55 switches on the transcription activation functions residing in the C-terminal effector domain. Here we report the crystal structure of N-Spo0A from Bacillus stearothermophilus at 1.6 A spacing, revealing a dimer formed by an alpha-helix swap. Comparison of this structure with the recently described structure of phosphorylated N-Spo0A shows that dimer formation results from a cis-trans isomerization of the Lys106--Pro107 peptide bond. The quaternary reorganization is associated with alterations in the active site stereochemistry which may have implications for signalling. Remarkably, this 3-D domain swapped N-Spo0A dimer has an identical topology to a hypothetical CheY-like dimer, recently proposed as an intermediate in the evolution of the family of periplasmic substrate binding proteins.
- Structural Biology Laboratory Department of Chemistry, University of York, York, YO10 5DD, UK.
Organizational Affiliation: