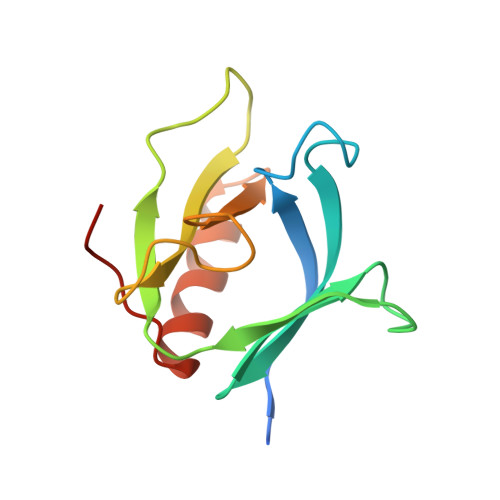
Crystal structure at 2.2 A resolution of the pleckstrin homology domain from human dynamin.
Ferguson, K.M., Lemmon, M.A., Schlessinger, J., Sigler, P.B.(1994) Cell 79: 199-209
- PubMed: 7954789 Search on PubMed
- DOI: https://doi.org/10.1016/0092-8674(94)90190-2
- Primary Citation Related Structures:
1DYN - PubMed Abstract:
The X-ray crystal structure of the pleckstrin homology (PH) domain from human dynamin has been refined to 2.2 A resolution. A seven-stranded beta sandwich of two orthogonal antiparallel beta sheets is closed at one corner by a C-terminal alpha helix. Opposite this helix are the three loops that vary most among PH domains. The basic fold is very similar to that of two other PH domains recently determined by nuclear magnetic resonance, confirming that PH domain with known structure is electrostatically polarized, with the three variable loops forming a positively charged surface. This surface includes the position of the X-linked immunodeficiency mutation in the Btk PH domain and may serve as a ligand-binding surface.
- Department of Molecular Biophysics and Biochemistry, Howard Hughes Medical Institute, Yale University, New Haven, Connecticut 06510.
Organizational Affiliation: