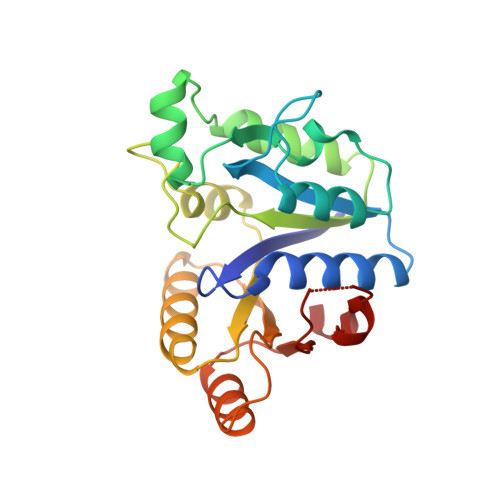
Crystal structure of an ATP-dependent carboxylase, dethiobiotin synthetase, at 1.65 A resolution.
Huang, W., Lindqvist, Y., Schneider, G., Gibson, K.J., Flint, D., Lorimer, G.(1994) Structure 2: 407-414
- PubMed: 8081756 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00042-3
- Primary Citation Related Structures:
1DTS - PubMed Abstract:
In Escherichia coli, the enzymes of the biotin biosynthesis pathway are encoded by the bio operon. One of these enzymes, ATP-dependent dethiobiotin synthetase, catalyzes the carboxylation of 7,8-diaminopelargonic acid leading to the formation of the ureido ring of biotin. The enzyme belongs to the class of ATP-dependent carboxylases and we present here the first crystal structure determined for this class of enzyme. We have determined the crystal structure of homodimeric dethiobiotin synthetase to 1.65 A resolution. The subunit consists of a seven-stranded parallel beta-sheet, surrounded by alpha-helices. The sheet contains the classical mononucleotide-binding motif with a fingerprint peptide Gly-X-X-X-X-X-Gly-Lys-Thr. The mononucleotide binding part of the structure is very similar to the GTP-binding protein H-ras-p21 and thus all GTP-binding proteins. A comparison reveals that some of the residues, which in H-ras-p21 interact with the nucleotide and the metal ion, are conserved in the synthetase. The three-dimensional structure of dethiobiotin synthetase has revealed that ATP-dependent carboxylases contain the classical mononucleotide-binding fold. Considerable similarities to the structure of the GTP-binding protein H-ras-p21 were found, indicating that both proteins might have evolved from a common ancestral mononucleotide-binding fold.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala.
Organizational Affiliation: