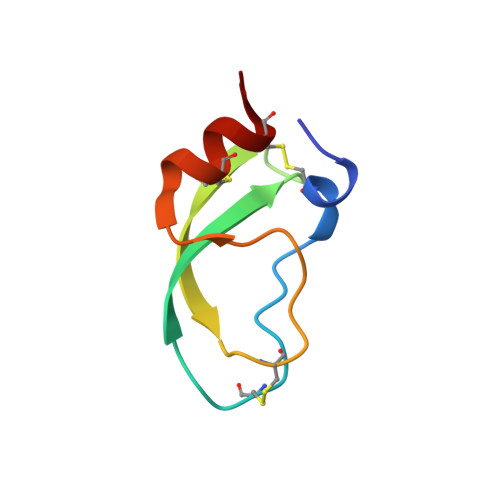
Nuclear magnetic resonance solution structure of dendrotoxin K from the venom of Dendroaspis polylepis polylepis.
Berndt, K.D., Guntert, P., Wuthrich, K.(1993) J Mol Biology 234: 735-750
- PubMed: 8254670 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1623
- Primary Citation Related Structures:
1DTK - PubMed Abstract:
The solution structure of dendrotoxin K (Toxin K), a protein consisting of one polypeptide chain with 57 residues purified from the venom of the black mamba, Dendroaspis polylepis polylepis, was determined by nuclear magnetic resonance (NMR) spectroscopy. On the basis of virtually complete sequence-specific 1H NMR assignments, including individual assignments for 38 pairs of diastereotopic substituents and side-chain amide protons, a total of 818 nuclear Overhauser effect distance constraints and 123 dihedral angle constraints were identified. Using this input, the solution structure of Toxin K was calculated with the program DIANA, and refined by restrained energy-minimization with a modified version of the program AMBER. The average root-mean-square deviation (r.m.s.d.) relative to the mean atomic co-ordinates of the 20 conformers selected to represent the solution structure is 0.31 A for all backbone atoms N, C alpha and C', and 0.90 A for all heavy-atoms of residues 2 to 56. The solution structure of Toxin K is very similar to the solution structure of the basic pancreatic trypsin inhibitor (BPTI) and the X-ray crystal structure of the alpha-dendrotoxin from Dendroaspis angusticeps (alpha-DTX), with r.m.s.d. values of 1.31 A and 0.92 A, respectively, for the backbone atoms of residues 2 to 56. Some local structural differences between Toxin K and BPTI are directly related to the fact that intermolecular interactions with two of the four internal molecules of hydration water in BPTI are replaced by intramolecular hydrogen bonds in Toxin K.
- Institut für Molekularbiologie und Biophysik Eidgenössische Technische Hochschule-Hönggerberg, Zürich, Switzerland.
Organizational Affiliation: