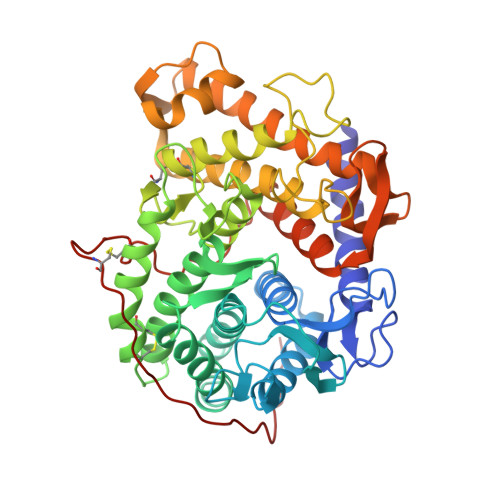
Refined structure for the complex of 1-deoxynojirimycin with glucoamylase from Aspergillus awamori var. X100 to 2.4-A resolution.
Harris, E.M., Aleshin, A.E., Firsov, L.M., Honzatko, R.B.(1993) Biochemistry 32: 1618-1626
- PubMed: 8431441 Search on PubMed
- DOI: https://doi.org/10.1021/bi00057a028
- Primary Citation Related Structures:
1DOG - PubMed Abstract:
The three-dimensional structure of the complex of 1-deoxynojirimycin with glucoamylase II-(471) from Aspergillus awamori var. X100 has been determined to 2.4-A resolution. The model includes residues corresponding to residues 1-471 of glucoamylase I from Aspergillus niger, two molecules of bound 1-deoxynojirimycin and 605 sites for water molecules. The crystallographic R factor from refinement is 0.119, and the root-mean-squared deviation in bond distances is 0.012 A. The inhibitor complex confirms the location of the active site in the packing void of the alpha/alpha-barrel as proposed by Aleshin et al. [Aleshin, A., Golubev, A., Firsov, L., & Honzatko, R. B. (1992) J. Biol. Chem. 267, 19291-19298]. One inhibitor molecule is associated with strong electron density and represents the principal site of interaction of 1-deoxynojirimycin with the enzyme. The other 1-deoxynojirimycin molecule is associated with weak electron density and therefore, probably represents a binding site of low affinity. Interactions of 1-deoxynojirimycin with the enzyme at its principal site involve Arg 45, Asp 55, Arg 305, and carbonyl 177. In addition, a water molecule (water 500) hydrogen bonds to Glu 400 and the 6-hydroxyl of 1-deoxynojirimycin and is at an approximate distance of 3.3 A from the "anomeric" carbon of the inhibitor. The structural arrangement of functional groups near the inhibitor molecule suggests that Glu 179 is a catalytic acid, Glu 400 a catalytic base, and water 500 the attacking nucleophile in the hydrolysis of maltooligosaccharides. The relevance of the X-ray work to proposed mechanisms of enzymatic hydrolysis of oligosaccharides is discussed.
- Department of Biochemistry and Biophysics, Iowa State University, Ames 50011.
Organizational Affiliation: