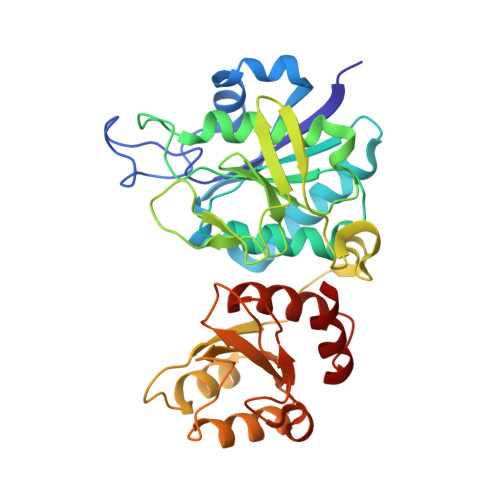
Reactions of Pseudomonas 7A glutaminase-asparaginase with diazo analogues of glutamine and asparagine result in unexpected covalent inhibitions and suggests an unusual catalytic triad Thr-Tyr-Glu.
Ortlund, E., Lacount, M.W., Lewinski, K., Lebioda, L.(2000) Biochemistry 39: 1199-1204
- PubMed: 10684596
- DOI: https://doi.org/10.1021/bi991797d
- Primary Citation of Related Structures:
1DJO, 1DJP - PubMed Abstract:
Pseudomonas 7A glutaminase-asparaginase (PGA) catalyzes the hydrolysis of D and L isomers of glutamine and asparagine. Crystals of PGA were reacted with diazo analogues of glutamine (6-diazo-5-oxo-L-norleucine, DON) and asparagine (5-diazo-4-oxo-L-norvaline, DONV), which are known inhibitors of the enzyme. The derivatized crystals remained isomorphous to native PGA crystals. Their structures were refined to crystallographic R = 0.20 and R(free) = 0.24 for PGA-DON and R = 0.19 and R = 0.23 for PGA-DONV. Difference Fourier electron density maps clearly showed that both DON and DONV inactivate PGA through covalent inhibition. Continuous electron density connecting the inhibitor to both Thr20 and Tyr34 of the flexible loop was observed providing strong evidence that Thr20 is the primary catalytic nucleophile and that Tyr34 plays an important role in catalysis as well. The unexpected covalent binding observed in the PGA-DON and PGA-DONV complexes shows that a secondary reaction involving the formation of a Tyr34-inhibitor bond takes place with concomitant inactivation of PGA. The predicted covalent linkage is not seen, however, suggesting an alternative method of inhibition not yet seen for these diazo analogues. These surprising results give insight as to the role of the flexible loop Thr and Tyr in the catalytic mechanism.
- Department of Chemistry and Biochemistry, University of South Carolina, Columbia, South Carolina 29208, USA.
Organizational Affiliation: