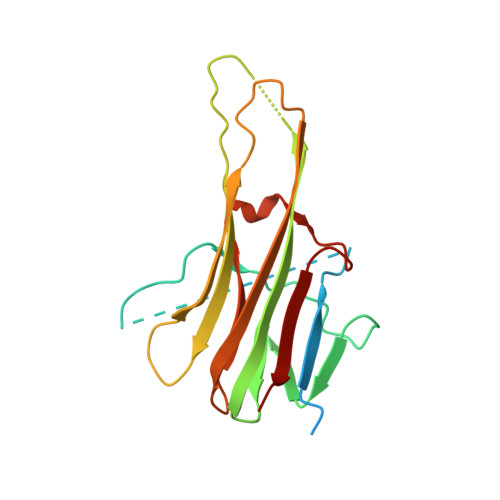
A unique zinc-binding site revealed by a high-resolution X-ray structure of homotrimeric Apo2L/TRAIL.
Hymowitz, S.G., O'Connell, M.P., Ultsch, M.H., Hurst, A., Totpal, K., Ashkenazi, A., de Vos, A.M., Kelley, R.F.(2000) Biochemistry 39: 633-650
- PubMed: 10651627 Search on PubMed
- DOI: https://doi.org/10.1021/bi992242l
- Primary Citation Related Structures:
1DG6 - PubMed Abstract:
Apoptosis-inducing ligand 2 (Apo2L, also called TRAIL), a member of the tumor necrosis factor (TNF) family, induces apoptosis in a variety of human tumor cell lines but not in normal cells [Wiley, S. R., Schooley, K., Smolak, P. J., Din, W. S., Huang, C.-P., Nicholl, J. K., Sutherland, G. R., Smith, T. D., Rauch, C., Smith, C. A., and Goodwin, R. G. (1995) Immunity 3, 673-682; Pitti, R. M., Marsters, S. A., Ruppert, S., Donahue, C. J., Moore, A., and Ashkenazi, A. (1996) J. Biol. Chem. 271, 12687-12690]. Here we describe the structure of Apo2L at 1.3 A resolution and use alanine-scanning mutagenesis to map the receptor contact regions. The structure reveals a homotrimeric protein that resembles TNF with receptor-binding epitopes at the interface between monomers. A zinc ion is buried at the trimer interface, coordinated by the single cysteine residue of each monomer. The zinc ion is required for maintaining the native structure and stability and, hence, the biological activity of Apo2L. This is the first example of metal-dependent oligomerization and function of a cytokine.
- Department of Protein Engineering, Genentech, Inc., 1 DNA Way, South San Francisco, California 94080, USA.
Organizational Affiliation: