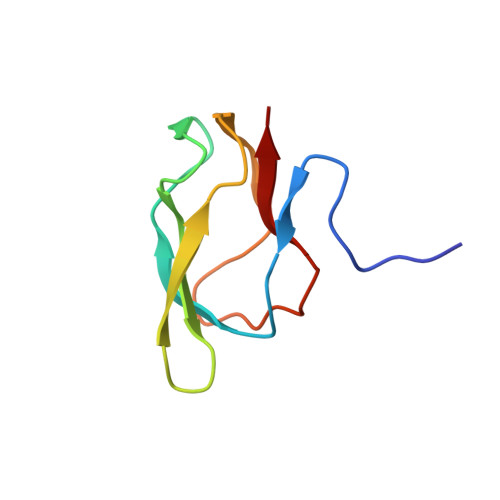
High resolution solution structure of the 1.3S subunit of transcarboxylase from Propionibacterium shermanii.
Reddy, D.V., Shenoy, B.C., Carey, P.R., Sonnichsen, F.D.(2000) Biochemistry 39: 2509-2516
- PubMed: 10704200 Search on PubMed
- DOI: https://doi.org/10.1021/bi9925367
- Primary Citation Related Structures:
1DCZ, 1DD2 - PubMed Abstract:
Transcarboxylase (TC) from Propionibacterium shermanii, a biotin-dependent enzyme, catalyzes the transfer of a carboxyl group from methylmalonyl-CoA to pyruvate to form propionyl-CoA and oxalacetate. Within the multi-subunit enzyme complex, the 1.3S subunit functions as the carboxyl group carrier and also binds the other two subunits to assist in the overall assembly of the enzyme. The 1.3S subunit is a 123 amino acid polypeptide (12.6 kDa) to which biotin is covalently attached at Lys 89. The three-dimensional solution structure of the full-length holo-1.3S subunit of TC has been solved by multidimensional heteronuclear NMR spectroscopy. The C-terminal half of the protein (51-123) is folded into a compact all-beta-domain comprising of two four-stranded antiparallel beta-sheets connected by short loops and turns. The fold exhibits a high 2-fold internal symmetry and is similar to that of the biotin carboxyl carrier protein (BCCP) of acetyl-CoA carboxylase, but lacks an extension that has been termed "protruding thumb" in BCCP. The first 50 residues, which have been shown to be involved in intersubunit interactions in the intact enzyme, appear to be disordered in the isolated 1.3S subunit. The molecular surface of the folded domain has two distinct surfaces: one side is highly charged, while the other comprises mainly hydrophobic, highly conserved residues.
- Department of Physiology & Biophysics, and the Department of Biochemistry, Case Western Reserve University, Cleveland, Ohio 44106, USA.
Organizational Affiliation: