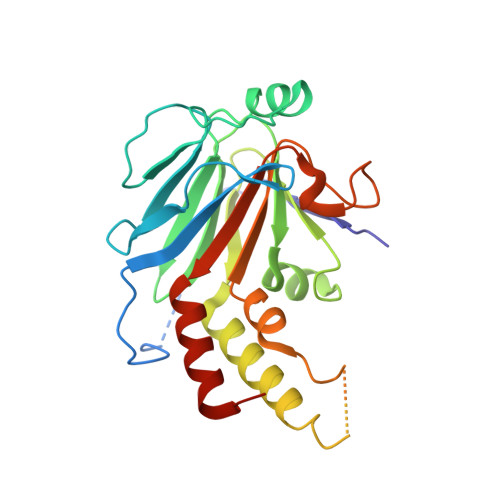
Crystal structure of a transcriptionally active Smad4 fragment.
Qin, B., Lam, S.S., Lin, K.(1999) Structure 7: 1493-1503
- PubMed: 10647180 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)88340-9
- Primary Citation Related Structures:
1DD1 - PubMed Abstract:
Smad4 functions as a common mediator of transforming growth factor beta (TGF-beta) signaling by forming complexes with the phosphorylated state of pathway-restricted SMAD proteins that act in specific signaling pathways to activate transcription. SMAD proteins comprise two domains, the MH1 and MH2 domain, separated by a linker region. The transcriptional activity and synergistic effect of Smad4 require a stretch of proline-rich sequence, the SMAD-activation domain (SAD), located N-terminal of the MH2 domain. To understand how the SAD contributes to Smad4 function, the crystal structure of a fragment including the SAD and MH2 domain (S4AF) was determined. The structure of the S4AF trimer reveals novel features important for Smad4 function. A Smad4-specific sequence insertion within the MH2 domain interacts with the C-terminal tail to form a structural extension from the core. This extension (the TOWER) contains a solvent-accessible glutamine-rich helix. The SAD reinforces the TOWER and the structural core through interactions; two residues involved in these interactions are targets of tumorigenic mutation. The solvent-accessible proline residues of the SAD are located on the same face as the glutamine-rich helix of the TOWER, forming a potential transcription activation surface. A tandem sulfate-ion-binding site was identified within the subunit interface, which may interact with the phosphorylated C-terminal sequence of pathway-restricted SMAD proteins. The structure suggests that the SAD provides transcriptional capability by reinforcing the structural core and coordinating with the TOWER to present the proline-rich and glutamine-rich surfaces for interaction with transcription partners. The sulfate-ion-binding sites are potential 'receptors' for the phosphorylated sequence of pathway-restricted SMAD proteins in forming a heteromeric complex. The structure thus provides a new model that can be tested using biochemical and cellular approaches.
- Department of Pharmacology and Molecular Toxicology, University of Massachusetts Medical School, Worcester 01655, USA.
Organizational Affiliation: