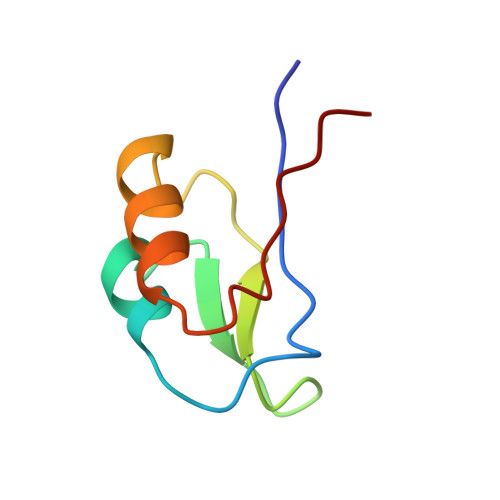
Determination of the structure of oxidised Desulfovibrio africanus ferredoxin I by 1H NMR spectroscopy and comparison of its solution structure with its crystal structure.
Davy, S.L., Osborne, M.J., Moore, G.R.(1998) J Mol Biology 277: 683-706
- PubMed: 9533888
- DOI: https://doi.org/10.1006/jmbi.1998.1631
- Primary Citation of Related Structures:
1DAX, 1DFD - PubMed Abstract:
The solution structure of the 64 amino acid Fe4S4 ferredoxin I from Desulfovibrio africanus has been determined using two-dimensional 1H NMR spectroscopy. Sequence-specific assignments were obtained for 59 amino acid residues and the structure determined with the program DIANA on the basis of 549 nuclear Overhauser enhancement (NOE) upper distance limits, and four dihedral angle and 52 distance constraints for the Fe4S4 cluster. The NMR structure was refined using the simulated annealing and energy minimisation protocols of the program X-PLOR to yield a final family of 19 structures selected on the basis of good covalent geometry and minimal restraint violations. The r.m.s.d. values to the average structure for this family are 0.49(+/-0.07) A and 0.94(+/-0.09) A for the backbone and heavy-atoms of residues 3 to 62, respectively. The NMR structure has been compared to the previously reported X-ray structures for the two molecules within the asymmetric unit of the crystal, which have a network of seven hydrogen bonds between them. This intermolecular interface, involving residues 38, 40 to 43 and 46, has the same conformation in the solution structures showing that the crystal packing does not perturb the structure. There are three regions in which the NMR and X-ray structures differ: around the cluster, a turn involving residues 8 to 10, and a loop involving residues 29 to 32. In the family of solution structures the backbone of the loop region incorporating residues 29 to 32 is well-defined whilst in both of the X-ray molecules it is ill-defined. The small differences between the X-ray and NMR structures for the cluster environment and the turn between residues 8 to 10 probably reflects a lack of NMR constraints. The observation of relatively rapid amide NH hydrogen exchange of NH groups close to the cluster, together with rapid flipping for Phe25, which is also close to the cluster, indicates that the cluster environment is more dynamic than the corresponding regions of related Fe/S proteins.
- School of Chemical Sciences, University of East Anglia, Norwich, NR4 7TJ, U.K.
Organizational Affiliation: