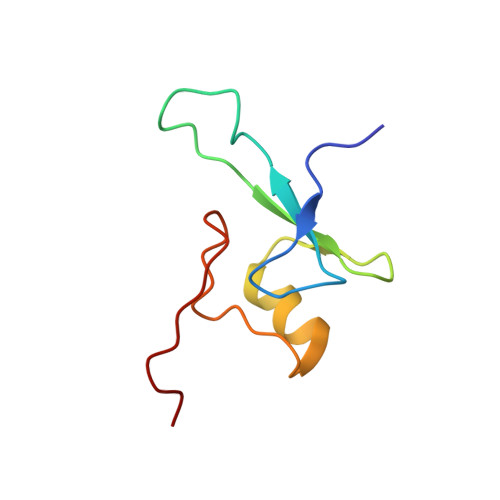
Solution structure of the methyl-CpG-binding domain of the methylation-dependent transcriptional repressor MBD1.
Ohki, I., Shimotake, N., Fujita, N., Nakao, M., Shirakawa, M.(1999) EMBO J 18: 6653-6661
- PubMed: 10581239 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.23.6653
- Primary Citation Related Structures:
1D9N - PubMed Abstract:
CpG methylation in vertebrates is important for gene silencing, alterations in chromatin structure and genomic stability, and differences in the DNA-methylation status are correlated with imprinting phenomena, carcinogenesis and embryonic development. Methylation signals are interpreted by protein factors that contain shared methyl-CpG-binding domains (MBDs). We have determined the solution structure of the MBD of the human methylation-dependent transcriptional repressor MBD1 by multi-dimensional heteronuclear NMR spectroscopy. It folds into an alpha/beta-sandwich structure with characteristic loops. Basic residues conserved in the MBD family are largely confined to one face of this fold and a flexible loop, which together form a large positively charged surface. Site-directed mutagenesis and chemical shift changes upon complexing with a methylated DNA facilitated identification of this surface as the DNA interaction site. In addition to three basic residues, conserved Tyr34 and Asp32 were shown to be important for the DNA binding.
- Graduate School of Biological Sciences, Nara Institute of Science and Technology, 8916-5 Takayama, Ikoma, Nara 630-0101, USA.
Organizational Affiliation: