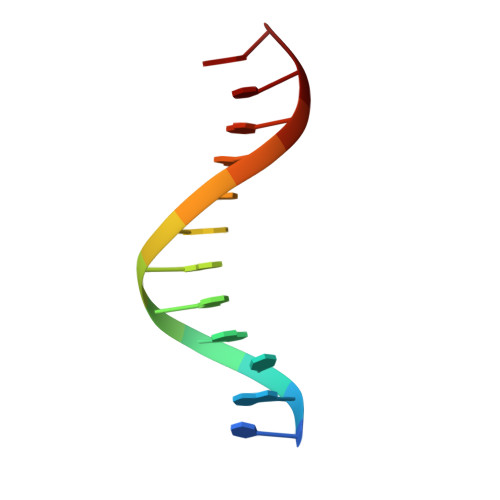
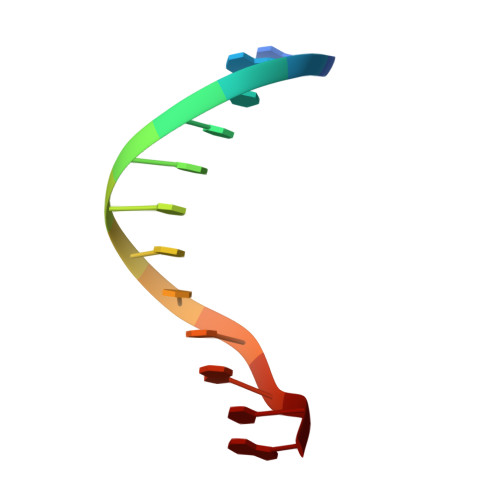
A DNA dodecamer containing an adenine tract crystallizes in a unique lattice and exhibits a new bend.
DiGabriele, A.D., Steitz, T.A.(1993) J Mol Biology 231: 1024-1039
- PubMed: 8515463
- DOI: https://doi.org/10.1006/jmbi.1993.1349
- Primary Citation Related Structures:
1D89 - PubMed Abstract:
The structure of d(CGCGAAAAAACG)/d(CGTTTTTTCGCG) was determined at 2.3 A resolution in order to deduce the local structural features that give rise to DNA bending by adenine tracts. Whereas all previously reported B-DNA dodecamers have crystallized isomorphously (spacegroup P2(1)2(1)2(1) with unit cell dimensions of a = 24.5 A, b = 40.3 A, c = 65.9 A), the duplex reported here crystallizes in a different lattice (spacegroup P2(1)2(1)2 with unit cell dimensions of a = 44.8 A, b = 66.1 A, c = 42.9 A). The DNA exhibits a 30 degree bend in the helix axis that is 180 degrees away from the 20 degree bend exhibited by the adenine tract DNA crystal structures that have been previously determined. This bend is 90 degrees away from the bend predicted for an adenine tract by solution and gel experiments. The adenine tract is straight and bending occurs in the G+C-rich regions. Comparison of the various adenine tract DNA crystal structures reveals that in all cases the adenine tracts have nearly identical structures, even though the overall bends of the helix axes are quite different. This implies that the structure of the adenine tract is robust, at least under the conditions of crystallization. The base-pairs in the adenine tracts exhibit a large propeller twist that leads to the formation of bifurcated hydrogen bonds and a narrow minor groove. In the crystal structure of d(CGCGAAAAAACG)/d(CGTTTTTTCGCG), a minor groove spine of hydration is observed that probably stabilizes the straight structure of the adenine tract. This straight structure of the A-tract is not consistent with the results of fiber diffraction, gel experiments, and NMR studies. Although this may imply that the results of solution experiments need to be reinterpreted, the conditions under which the crystals were grown are different from those under which the solution experiments were done. The possibility remains that the 2-methyl-2,4-pentanediol necessary for crystal growth may facilitate formation of the spine of hydration that stabilizes the straight A-tract, although recent NMR results show the presence of the spine of hydration in aqueous solution. We have also extended our previously reported observation that non-self-complementary DNA structures can exist in the crystal lattice in two orientations. A dodecamer brominated on one strand provides experimental evidence that d(CGCAAAAAAGCG)/d(CGCTTTTTTG CG) is positioned in two orientations in the crystal lattice that are related by a 180 degree rotation around the pseudo-dyad axis of the sequence.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Chemistry, Yale University, New Haven, CT 06511.
Organizational Affiliation: