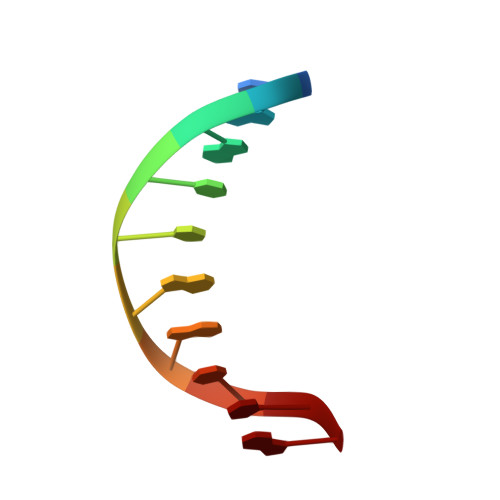
Structure of a B-DNA decamer with a central T-A step: C-G-A-T-T-A-A-T-C-G.
Quintana, J.R., Grzeskowiak, K., Yanagi, K., Dickerson, R.E.(1992) J Mol Biology 225: 379-395
- PubMed: 1593626 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90928-d
- Primary Citation Related Structures:
1D49 - PubMed Abstract:
The X-ray crystal structure analysis of the decamer C-G-A-T-T-A-A-T-C-G has been carried out to a resolution of 1.5 A. The crystals are space group P2(1)2(1)2(1), cell dimensions a = 38.60 A, b = 39.10 A, c = 33.07 A. The structure was solved by molecular replacement and refined with X-PLOR and NUCLSQ. The final R factor for a model with 404 DNA atoms, 108 water molecules and one magnesium hexahydrate cation is 15.7%. The double helix is essentially isostructural with C-G-A-T-C-G-A-T-C-G, with closely similar local helix parameters. The structure of the T-T-A-A center differs from that found in C-G-C-G-T-T-A-A-C-G-C-G in that the minor groove in our decamer is wide at the central T-A step rather than narrow, and the twist angle of the T-A step is small (31.1 degrees) rather than large. Whereas the tetrad model provides a convenient framework for discussing local DNA helix structure, it cannot be the entire story. The articulated helix model of DNA structure proposes that certain sequence regions of DNA show preferential twisting or bending properties, whereas other regions are less capable of deformation, in a manner that may be useful in sequence recognition by drugs and protein. Further crystal structure analyses should help to delineate the precise nature of sequence-dependent articulation in the DNA double helix.
- Department of Chemistry and Biochemistry, University of California, Los Angeles 90024.
Organizational Affiliation: