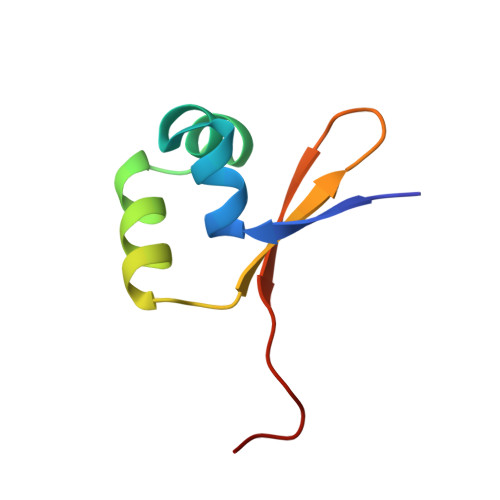
The structural basis for enhanced stability and reduced DNA binding seen in engineered second-generation Cro monomers and dimers.
Rupert, P.B., Mollah, A.K., Mossing, M.C., Matthews, B.W.(2000) J Mol Biology 296: 1079-1090
- PubMed: 10686105 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.3498
- Primary Citation Related Structures:
1D1L, 1D1M - PubMed Abstract:
It was previously shown that the Cro repressor from phage lambda, which is a dimer, can be converted into a stable monomer by a five-amino acid insertion. Phe58 is the key residue involved in this transition, switching from interactions which stabilize the dimer to those which stabilize the monomer. Structural studies, however, suggested that Phe58 did not penetrate into the core of the monomer as well as it did into the native dimer. This was strongly supported by the finding that certain core-repacking mutations, including in particular, Phe58-->Trp, increased the stability of the monomer. Unexpectedly, the same substitution also increased the stability of the native dimer. At the same time it decreased the affinity of the dimer for operator DNA. Here we describe the crystal structures of the Cro F58W mutant, both as the monomer and as the dimer. The F58W monomer crystallized in a form different from that of the original monomer. In contrast to that structure, which resembled the DNA-bound form of Cro, the F58W monomer is closer in structure to wild-type (i.e. non-bound) Cro. The F58W dimer also crystallizes in a form different from the native dimer but has a remarkably similar overall structure which tends to confirm the large changes in conformation of Cro on binding DNA. Introduction of Trp58 perturbs the position occupied by the side-chain of Arg38, a DNA-contact residue, providing a structural explanation for the reduction in DNA-binding affinity. The improved thermal stability is seen to be due to the enhanced solvent transfer free energy of Trp58 relative to Phe58, supplemented in the dimer structure, although not the monomer, by a reduction in volume of internal cavities.
- Institute of Molecular Biology Howard Hughes Medical Institute and Department of Physics 1229, University of Oregon, Eugene, OR 97403, USA.
Organizational Affiliation: