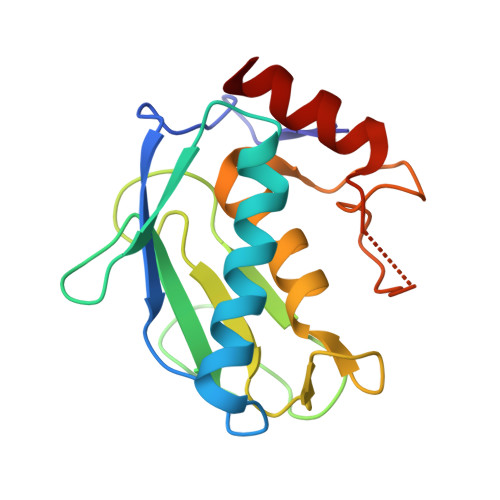
Structure of recombinant mouse collagenase-3 (MMP-13).
Botos, I., Meyer, E., Swanson, S.M., Lemaitre, V., Eeckhout, Y., Meyer, E.F.(1999) J Mol Biology 292: 837-844
- PubMed: 10525409
- DOI: https://doi.org/10.1006/jmbi.1999.3068
- Primary Citation Related Structures:
1CXV - PubMed Abstract:
The matrix metalloproteinases are crucial in the physiological and pathological degradation of the mammalian extracellular matrix, including breast tumours, and osteoarthritic cartilage. These enzymes are classified according to their matrix substrate specificity. Collagenase-3 (MMP-13) is a member of this family and preferentially cleaves type II collagen, cartilage, fibronectin and aggrecan. Collagenase-3 is normally expressed in hypertrophic chondrocytes, periosteal cells, and osteoblasts during bone development. The structure of the catalytic domain of recombinant mouse collagenase-3, complexed to the hydroxamate inhibitor (RS-113456), is reported at 2.0 A resolution. Molecular replacement and weak phasing information from a single derivative determined the structure. Neither molecular replacement nor derivative methods had a sufficient radius of convergence to yield a refinable structure. The structure illuminates the atomic zinc ion interactions with functional groups in the active site, emphasizing zinc ligation and the very voluminous hydrophobic P1' group for the inhibitor potency. The structure provides insight into the specificity of this enzyme, facilitating design of specific inhibitors to target various diseases.
- Department of Biochemistry and Biophysics, Texas A&M University, TX, 77843-2128, USA.
Organizational Affiliation: