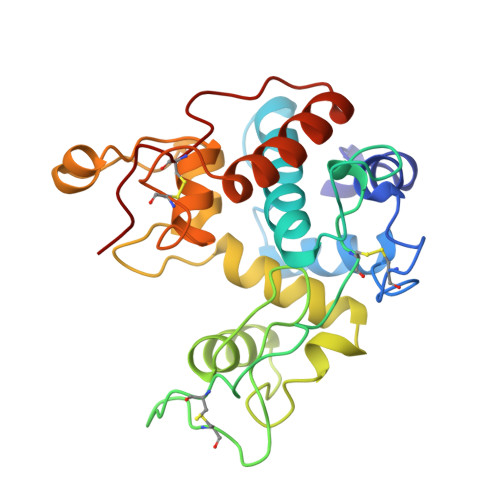
Refined structure of the chitinase from barley seeds at 2.0 a resolution.
Song, H.K., Suh, S.W.(1996) Acta Crystallogr D Biol Crystallogr 52: 289-298
- PubMed: 15299702 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444995009061
- Primary Citation Related Structures:
1CNS - PubMed Abstract:
Chitinase from barley seeds is a monomeric enzyme with 243 amino-acid residues and it plays a role as a defense protein. Its structure, previously determined at 2.8 A resolution by multiple isomorphous replacement method, is mainly alpha-helical [Hart, Monzingo, Ready, Ernst & Robertus, (1993). J. Mol. Biol. 229, 189-193]. The crystallization and preliminary X-ray data of the same enzyme in a different crystal form has been reported independently [Song, Hwang, Kim & Suh, (1993). Proteins, 17, 107-109}, the asymmetric unit of which contains two chitinase molecules. As a step toward understanding the general principles of catalysis, reported here is the structure of chitinase from barley seeds in this crystal form, as determined by molecular replacement and subsequently refined at 2.0 A resolution, with incorporation of partial data to 1.9 A (R factor of 18.9% for 31 038 unique reflections with F(o)> 2sigma(F) in the range 8.0-1.9 A). The r.m.s. deviations from ideal stereochemistry are 0.013 A for bond lengths and 1.32 degrees for bond angles. A superposition of the two independent molecules in the asymmetric unit gives an r.m.s. difference of 0.55 A for all protein atoms (0.43 and 0.74 A for main-chain and side-chain atoms, respectively). When the refined model of each chitinase molecule in the asymmetric unit is superposed with the starting model, the r.m.s. difference for all shared protein atoms is 0.99 A for molecule 1 and 0.85 A for molecule 2, respectively. Through a sequence comparison with homologous plant chitinases as well as a structural comparison with the active sites of other glycosidases, key catalytic residues have been identified and the active site has been located in the three-dimensional structure of the barley chitinase. The present structure, refined at an effective resolution of 2.0 A with incorporation of partial data to 1.9 A, represents a significant improvement in resolution compared to the previously reported model. The improved resolution has enabled the location of solvent atoms, including water molecules near the catalytic residues, in addition to the positioning of protein atoms with greater accuracy.
- Department of Chemistry and Center for Molecular Caralysis, Seoul National University, Korea.
Organizational Affiliation: