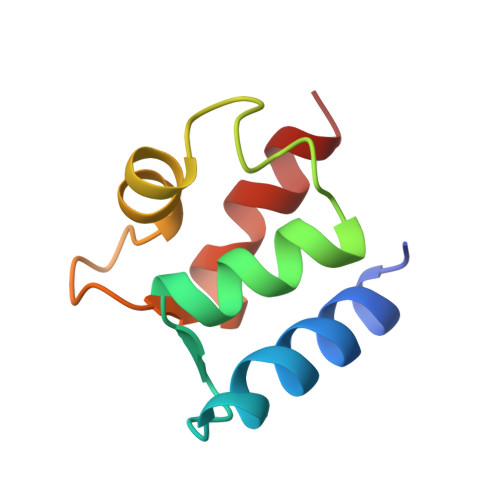
Solution structure of (Cd2+)1-calbindin D9k reveals details of the stepwise structural changes along the Apo-->(Ca2+)II1-->(Ca2+)I,II2 binding pathway.
Akke, M., Forsen, S., Chazin, W.J.(1995) J Mol Biology 252: 102-121
- PubMed: 7666423
- DOI: https://doi.org/10.1006/jmbi.1995.0478
- Primary Citation Related Structures:
1CDN - PubMed Abstract:
The three-dimensional solution structure of (Cd2+)1-calbindin D9k has been determined by distance geometry, restrained molecular dynamics and relaxation matrix calculations using experimental constraints obtained from two-dimensional 1H and 15N-1H NMR spectroscopy. The final input data consisted of 1055 NOE distance constraints and 71 dihedral angle constraints, corresponding to 15 constraints per residue on average. The resulting ensemble of 24 structures has no distance or dihedral angle constraints consistently violated by more than 0.07 A and 1.8 degrees, respectively. The structure is characteristic of an EF-hand protein, with two helix-loop-helix calcium binding motifs joined by a flexible linker, and a short anti-parallel beta-type interaction between the two ion-binding sites. The four helices are well defined with a root mean square deviation from the mean coordinates of 0.35 A for the backbone atoms. The structure of the half-saturated cadmium state was compared with the previously determined solution structures of the apo and fully calcium saturated calbindin D9k. The comparisons were aided by introducing the ensemble averaged distance difference matrix as a tool for analyzing differences between two ensembles of structures. Detailed analyses of differences between the three states in backbone and side-chain dihedral angles, hydrogen bonds, interatomic distances, and packing of the hydrophobic core reveal the reorganization of the protein that occurs upon ion binding. Overall, it was found that (Cd2+)1-calbindin D9k, representing the half-saturated calcium state with an ion in site II, is structurally more similar to the fully calcium-saturated state than the apo state. Thus, for the binding sequence apo-->(Ca2+)II1-->(Ca2+)I,II2, the structural changes occurring upon ion binding are most pronounced for the first binding step, an observation that bears significantly on the molecular basis for cooperative calcium binding in calbindin D9k.
- Department of Molecular Biology, Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: