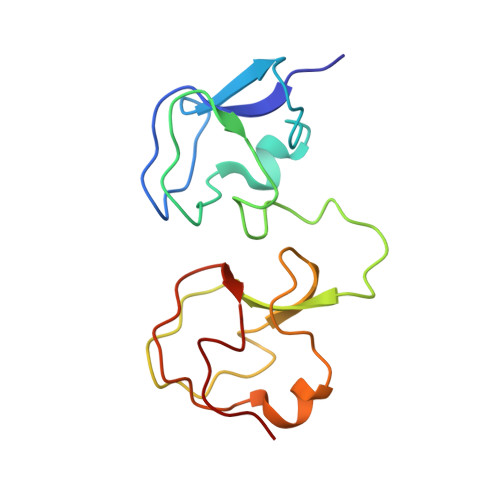
NMR analysis of type III antifreeze protein intramolecular dimer. Structural basis for enhanced activity.
Miura, K., Ohgiya, S., Hoshino, T., Nemoto, N., Suetake, T., Miura, A., Spyracopoulos, L., Kondo, H., Tsuda, S.(2001) J Biological Chem 276: 1304-1310
- PubMed: 11010977
- DOI: https://doi.org/10.1074/jbc.M007902200
- Primary Citation of Related Structures:
1C89, 1C8A - PubMed Abstract:
The structure of a new antifreeze protein (AFP) variant, RD3, from antarctic eel pout (Rhigophila dearborni) with enhanced activity has been determined for the first time by nuclear magnetic resonance spectroscopy. RD3 comprises a unique translational topology of two homologous type III AFP globular domains, each containing one flat, ice binding plane. The ice binding plane of the N domain is located approximately 3.5 A "behind" that of the C domain. The two ice binding planes are located laterally with an angle of 32 +/- 12 degrees between the planes. These results suggest that the C domain plane of RD3 binds first to the ice [1010] prism plane in the <0001> direction, which induces successive ice binding of the N domain in the <0101> direction. This manner of ice binding caused by the unique structural topology of RD3 is thought to be crucial for the significant enhancement of antifreeze activity, especially at low AFP concentrations.
- Bioscience and Chemistry Division, Hokkaido National Industrial Research Institute, 2-17-2-1 Tsukisamu-Higashi, Toyohira, Sapporo 062-8517, Japan.
Organizational Affiliation: