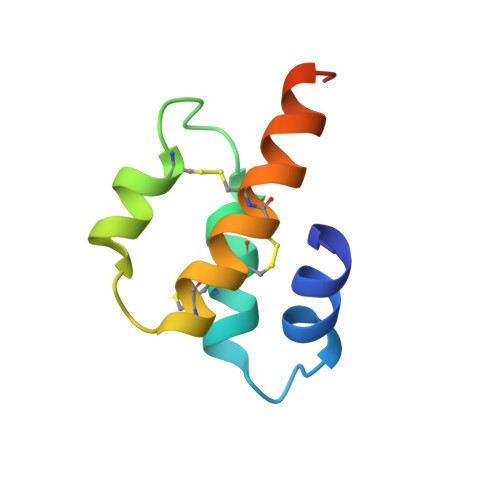
Three-dimensional structure of porcine C5adesArg from 1H nuclear magnetic resonance data.
Williamson, M.P., Madison, V.S.(1990) Biochemistry 29: 2895-2905
- PubMed: 2337573
- DOI: https://doi.org/10.1021/bi00464a002
- Primary Citation of Related Structures:
1C5A - PubMed Abstract:
Two-dimensional nuclear magnetic resonance spectra of porcine C5adesArg (73 residues) have been used to construct a list of 34 hydrogen bonds, 27 dihedral angle constraints, and 151 distance constraints, derived from nuclear Overhauser effect data. These constraints were used in restrained molecular dynamics calculations on residues 1-65 of C5a, starting from a folded structure modeled on the crystal structure of a homologous protein, C3a. Forty-one structures have been calculated, which fall into three similar families with few violations of the imposed constraints. Structures in the most populated family have a root-mean-square deviation from the average structure of 1.02 A for the C alpha atoms, with good definition of the internal residues. There is good agreement between the calculated structures and other nuclear magnetic resonance data. The structure is very similar to that recently reported for human C5a [Zuiderweg et al. (1989) Biochemistry 28, 172-185]. Some biological implications of these structures are discussed.
- Physical Methods Department, Roche Products Ltd., Welwyn Garden City, Herts, U.K.
Organizational Affiliation: