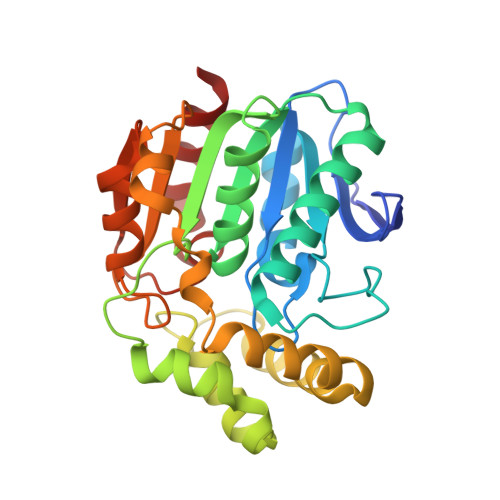
Three-Dimensional Structure of Microbial 2-Hydroxyl-6-Oxo-6-Phenylhexa-2,4- Dienoic Acid (Hpda) Hydrolase (Bphd Enzyme) from Rhodococcus Sp. Strain Rha1, in the Pcb Degradation Pathway
Nandhagopal, N., Senda, T., Hatta, T., Yamada, A., Masai, E., Fukuda, M., Mitsui, Y.(1997) Proc Jpn Acad Ser B Phys Biol Sci 73: 154