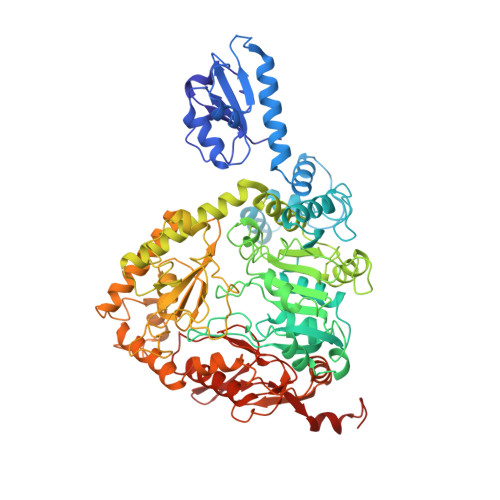
Three-dimensional structure of the Gly121Tyr dimeric form of ornithine decarboxylase from Lactobacillus 30a.
Vitali, J., Carroll, D., Chaudhry, R.G., Hackert, M.L.(1999) Acta Crystallogr D Biol Crystallogr 55: 1978-1985
- PubMed: 10666573
- DOI: https://doi.org/10.1107/s0907444999010756
- Primary Citation of Related Structures:
1C4K - PubMed Abstract:
Ornithine decarboxylases catalyze the conversion of ornithine to putrescine at the beginning of the polyamine pathway. Ornithine decarboxylase (ODC) from Lactobacillus 30a is a 990612 Da dodecamer composed of six homodimers. A single point mutation (Gly121Tyr) was found to prevent association of dimers into dodecamers. The dimeric protein has been crystallized at pH 7.0 in the presence of guanosine triphosphate (GTP). Crystals belong to space group P3(2)21, with unit-cell parameters a = 111.8, c = 135.9 A and one monomer in the asymmetric unit. The structure was determined by molecular replacement and refined using simulated annealing to R = 0.211 at 2. 7 A resolution. The GTP-binding site was analyzed in detail. The protein exhibits a novel binding mode for GTP which is different from that seen in most G-proteins or GTPases. Central to this binding scheme appear to be three lysines, Lys190, Lys374 and Lys382, which form salt bridges with the three phosphates, and Thr191, which hydrogen bonds with the guanine base. Furthermore, the structure suggests that there is some flexibility in the wing domain, which can change its orientation as the protein adapts to its environment. The active site is similar to that of the native enzyme, consistent with the observation that the enzyme activity does not depend on its dodecameric state.
- Department of Chemistry and Biochemistry, University of Texas, Austin, TX 78712, USA.
Organizational Affiliation: