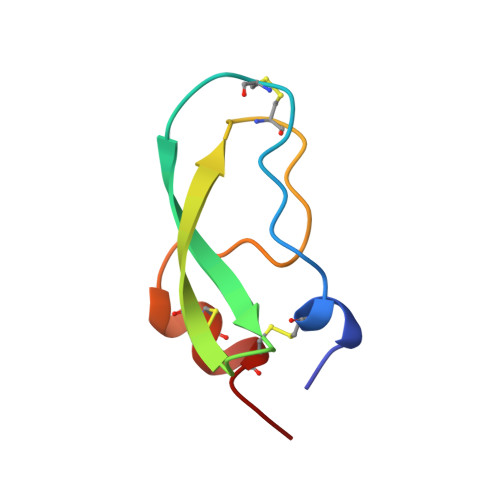
The crystal structure of anionic salmon trypsin in complex with bovine pancreatic trypsin inhibitor.
Helland, R., Leiros, I., Berglund, G.I., Willassen, N.P., Smalas, A.O.(1998) Eur J Biochem 256: 317-324
- PubMed: 9760170 Search on PubMed
- DOI: https://doi.org/10.1046/j.1432-1327.1998.2560317.x
- Primary Citation Related Structures:
1BZX - PubMed Abstract:
The complex formed between anionic salmon trypsin (ST) and bovine pancreatic trypsin inhibitor (BPTI) has been crystallised, and the X-ray structure has been solved using the molecular replacement method. The crystals are hexagonal and belong to space group P6(1)22 with lattice parameters of a = b = 83.12 A and c = 222.15 A. Data have been collected to 2.1 A and the structure has been refined to a crystallographic R-factor of 20.6%. Catalysis by salmon trypsin is distinguished by a Km value 20-fold lower than that for mammalian trypsins, and a k(cat) twice as high. The present ST-BPTI complex serves as a model for the Michaelis-Menten complex, and has been compared with corresponding bovine and rat trypsin (RT) complexes. The binding of BPTI to salmon trypsin is characterised by stronger primary interactions in the active site, and a somewhat looser secondary binding.
- Department of Chemistry, University of Tromsø, Norway.
Organizational Affiliation: