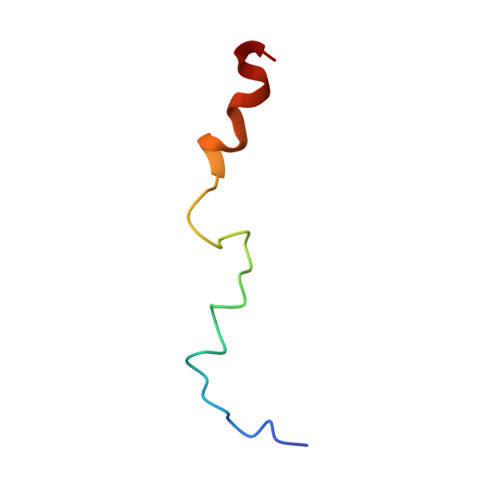
Structure of a biologically active fragment of human serum apolipoprotein C-II in the presence of sodium dodecyl sulfate and dodecylphosphocholine.
Storjohann, R., Rozek, A., Sparrow, J.T., Cushley, R.J.(2000) Biochim Biophys Acta 1486: 253-264
- PubMed: 10903476 Search on PubMed
- DOI: https://doi.org/10.1016/s1388-1981(00)00062-7
- Primary Citation Related Structures:
1BY6 - PubMed Abstract:
We have studied the three-dimensional structure of a biologically active peptide of apolipoprotein C-II (apoC-II) in the presence of lipid mimetics by CD and NMR spectroscopy. This peptide, corresponding to residues 44-79 of apoC-II, has been shown to reverse the symptoms of genetic apoC-II deficiency in a human subject. A comparison of alpha-proton secondary shifts and CD spectroscopic data indicates that the structure of apoC-II(44-79) is similar in the presence of dodecylphosphocholine and sodium dodecyl sulfate. The three-dimensional structure of apoC-II(44-79) in the presence of sodium dodecyl sulfate, determined by relaxation matrix calculations, contains two amphipathic helical domains formed by residues 50-58 and 67-75, separated by a non-helical linker centered at Tyr63. The C-terminal helix is terminated by a loop formed by residues 76-79. The C-terminal helix is better defined and has a larger hydrophobic face than the N-terminal helix, which leads us to propose that the C-terminal helix together with the non-helical Ile66 constitute the primary lipid binding domain of apoC-II(44-79). Based on our structure we suggest a new mechanism of lipoprotein lipase activation in which both helices of apoC-II(44-79) remain lipid bound, while the seven-residue interhelical linker extends away from the lipid surface in order to project Tyr63 into the apoC-II binding site of lipoprotein lipase.
- Institute of Molecular Biology and Biochemistry and Department of Chemistry, Simon Fraser University, Burnaby, British Columbia, Canada.
Organizational Affiliation: