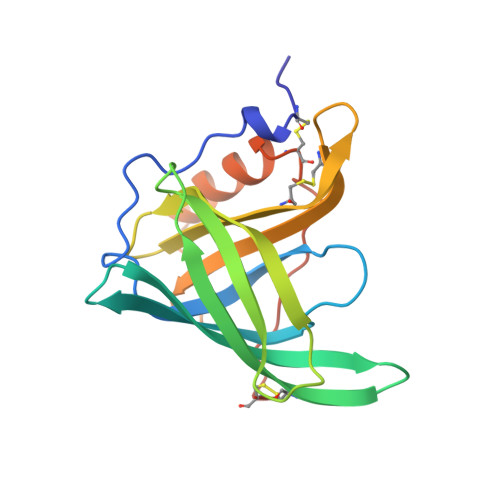
Crystal structure of the trigonal form of human plasma retinol-binding protein at 2.5 A resolution.
Zanotti, G., Ottonello, S., Berni, R., Monaco, H.L.(1993) J Mol Biology 230: 613-624
- PubMed: 8464067 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1173
- Primary Citation Related Structures:
1BRP, 1BRQ - PubMed Abstract:
The three-dimensional structures of the liganded and unliganded forms of human plasma retinol binding protein (RBP) in the trigonal crystal form have been solved at 2.5 A resolution. The final model of RBP complexed with retinol (holoRBP, space group R3, a = b = 104.0 A, c = 74.4 A) has a crystallographic R factor of 0.176 for 9652 reflections. The unliganded form, obtained through a purification procedure which included steps based on hydrophobic interaction chromatography, crystallized isomorphously with holoRBP and its structure has been refined to an R factor of 0.190 for 9614 reflections. The structure of the trigonal holo protein is quite similar to that of the orthorhombic form: the root-mean-square deviation of all the equivalent alpha-carbons in the two chains is 0.53 A. The structural comparison between the liganded and unliganded forms of RBP in the crystal did not reveal gross conformational changes. The most significant difference between the two forms of the protein is a conformational change involving residues from 34 to 37. In this region, the movements of side-chains of Leu35 and Phe36 are most noticeable. In particular, in the unliganded form the side-chain ring of the latter residue is in the place previously occupied by the alcoholic moiety of retinol. Our data are consistent with a model in which a region comprising these residues and at least part of the opening of the beta-barrel is involved in the recognition between RBP and transthyretin. In the case of the unliganded form, the central cavity, that is occupied by the vitamin in the two human crystalline holoRBPs, is filled by electron density that, at the present resolution, we interpret as solvent.
- Department of Organic Chemistry, University of Padova, Italy.
Organizational Affiliation: