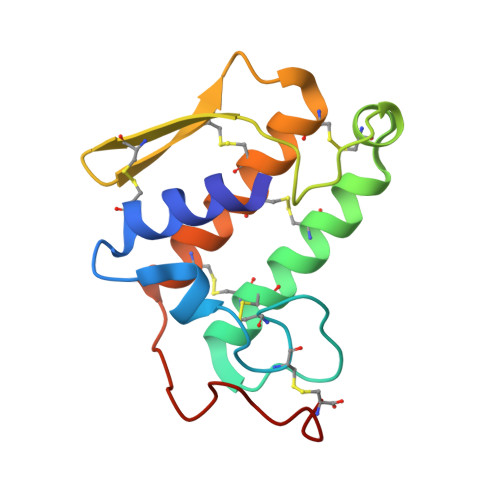
Phospholipase A2 engineering. X-ray structural and functional evidence for the interaction of lysine-56 with substrates.
Noel, J.P., Bingman, C.A., Deng, T.L., Dupureur, C.M., Hamilton, K.J., Jiang, R.T., Kwak, J.G., Sekharudu, C., Sundaralingam, M., Tsai, M.D.(1991) Biochemistry 30: 11801-11811
- PubMed: 1751497 Search on PubMed
- DOI: https://doi.org/10.1021/bi00115a010
- Primary Citation Related Structures:
1BPQ, 2BPP - PubMed Abstract:
Site-directed mutagenesis studies of bovine pancreatic phospholipase A2 (PLA2, overproduced in Escherichia coli) showed that replacement of surface residue Lys-56 by a neutral or hydrophobic amino acid residue resulted in an unexpected and significant change in the function of the enzyme. The kcat for phosphatidylcholine micelles increases 3-4-fold for K56M, K56I, and K56F and ca. 2-fold for K56N and K56T but does not change for K56R. These results suggest that the side chain of residue 56 has significant influence on the activity of PLA2. In order to probe the structural basis for the enhanced activity, the crystal structures of wild-type and K56M PLA2 were determined by X-ray crystallography to a resolution of 1.8 A. The results suggest that the mutation has not only perturbed the conformation of the side chain of Met-56 locally but also caused conformational changes in the neighboring loop (residues 60-70), resulting in the formation of a hydrophobic pocket by residues Met-56, Tyr-52, and Tyr-69. Docking of a phosphatidylcholine inhibitor analogue into the active site of K56M, according to the structure of the complex of cobra venom PLA2-phosphatidylethanolamine inhibitor analogue [White, S.P., Scott, D. L., Otwinowski, Z., Gleb, M. H., & Sigler, P. (1990) Science 250, 1560-1563], showed that the choline moiety [N(CH3)3]+ is readily accommodated into the newly formed hydrophobic pocket with a high degree of surface complementarity. This suggests a possible interaction between residue 56 and the head group of the phospholipid, explaining the enhanced activities observed when the positively charged Lys-56 is substituted by apolar residues, viz., K56M, K56I, and K56F. Further support for this interpretation comes from the 5-fold enhancement in kcat for the mutant K56E with a negatively charged side chain, where there would be an attractive electrostatic interaction between the side chain of Glu-56 and the positively charged choline moiety. Our results also refute a recent report [Tomasselli, A. G., Hui, J., Fisher, J., Zürcher-Neely, H., Reardon, I.M., Oriaku, E., Kézdy, F.J., & Heinrikson, R.L. (1989) J. Biol. Chem. 264, 10041-10047] that substrate-level acylation of Lys-56 is an obligatory step in the catalysis by PLA2.
- Department of Chemistry, Ohio State University, Columbus 43210.
Organizational Affiliation: