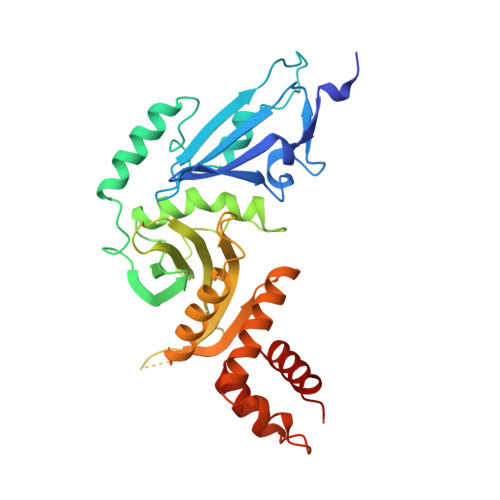
Structure of the histone acetyltransferase Hat1: a paradigm for the GCN5-related N-acetyltransferase superfamily.
Dutnall, R.N., Tafrov, S.T., Sternglanz, R., Ramakrishnan, V.(1998) Cell 94: 427-438
- PubMed: 9727486
- DOI: https://doi.org/10.1016/s0092-8674(00)81584-6
- Primary Citation of Related Structures:
1BOB - PubMed Abstract:
We have solved the crystal structure of the yeast histone acetyltransferase Hat1-acetyl coenzyme A (AcCoA) complex at 2.3 A resolution. Hat1 has an elongated, curved structure, and the AcCoA molecule is bound in a cleft on the concave surface of the protein, marking the active site of the enzyme. A channel of variable width and depth that runs across the protein is probably the binding site for the histone substrate. A model for histone H4 binding by Hat1 is discussed in terms of possible sources of specific lysine recognition by the enzyme. The structure of Hat1 provides a model for the structures of the catalytic domains of a protein superfamily that includes other histone acetyltransferases such as Gcn5 and CBP.
- Department of Biochemistry, University of Utah School of Medicine, Salt Lake City 84132, USA. rnd@snowbird.med.utah.edu
Organizational Affiliation: