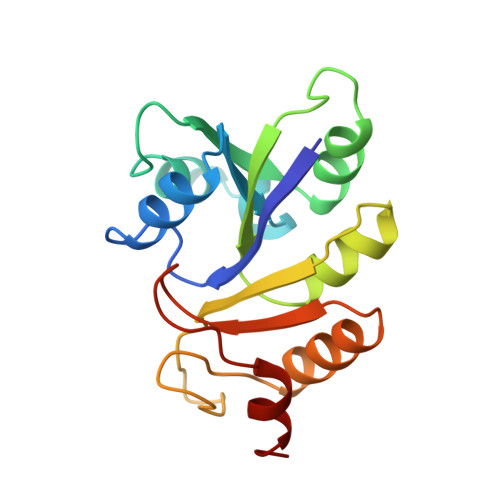
Crystal structure of the IIB subunit of a fructose permease (IIBLev) from Bacillus subtilis.
Schauder, S., Nunn, R.S., Lanz, R., Erni, B., Schirmer, T.(1998) J Mol Biology 276: 591-602
- PubMed: 9551099
- DOI: https://doi.org/10.1006/jmbi.1997.1544
- Primary Citation of Related Structures:
1BLE - PubMed Abstract:
The bacterial phosphoenolpyruvate-dependent phosphotransferase system (PTS) mediates both the uptake of carbohydrates across the cytoplasmic membrane and their phosphorylation. During this process, a phosphoryl group is transferred from phosphoenolpyruvate via the general PTS proteins enzyme I, HPr and the sugar-specific components IIA, IIB to the transported sugar. The crystal structure of the IIB subunit of a fructose transporter from Bacillus subtilis (IIBLev) was solved by MIRAS to a resolution of 2.9 A. IIBLev comprises 163 amino acid residues that are folded into an open, mainly parallel beta-sheet with helices packed on either face. The phosphorylation site (His15) is located on the first loop (1/A) at one of the topological switch-points of the fold. Despite different global folds, IIBLev and HPr have very similar active-site loop conformations with the active-site histidine residues located close to the N terminus of the first helix. This resemblance may be of functional importance, since both proteins exchange a phosphoryl group with the same IIA subunit. The structural basis of phosphoryl transfer from HPr to IIAMan to IIBMan was investigated by modeling of the respective transition state complexes using the known HPr and IIAMan structures and a homology model of IIBMan that was derived from the IIBLev structure. All three proteins contain a helix that appears to be suitable for stabilization of the phospho-histidine by dipole and H-bonding interactions. Smooth phosphoryl transfer from one N-cap position to the other appears feasible with a minimized transition state energy due to simultaneous interactions with the donor and the acceptor helix.
- Department of Structural Biology, University of Basel, Switzerland.
Organizational Affiliation: