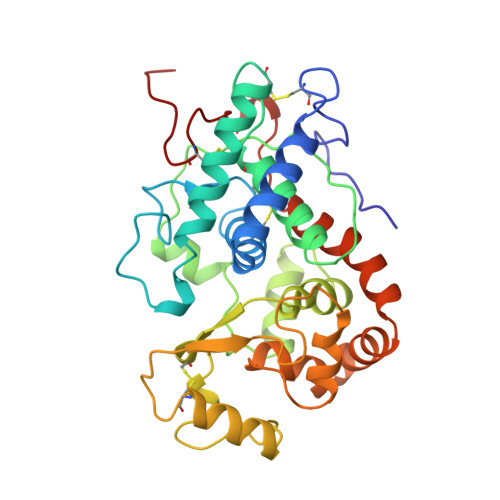
Structure of barley grain peroxidase refined at 1.9-A resolution. A plant peroxidase reversibly inactivated at neutral pH.
Henriksen, A., Welinder, K.G., Gajhede, M.(1998) J Biological Chem 273: 2241-2248
- PubMed: 9442067
- DOI: https://doi.org/10.1074/jbc.273.4.2241
- Primary Citation of Related Structures:
1BGP - PubMed Abstract:
The crystal structure of the major peroxidase of barley grain (BP 1) has been solved by molecular replacement and phase combination and refined to an R-factor of 19.2% for all data between 38 and 1.9 A. The refined model includes amino acid residues 1-309, one calcium ion, one sodium ion, iron-protoporphyrin IX, and 146 solvent molecules. BP 1 has the apparently unique property of being unable to catalyze the reaction with the primary substrate hydrogen peroxide to form compound I at pH values > 5, a feature investigated by obtaining crystal structure data at pH 5.5, 7.5, and 8.5. Structural comparison shows that the overall fold of inactive barley grain peroxidase at these pH values resembles that of both horseradish peroxidase C and peanut peroxidase. The key differences between the structures of active horseradish peroxidase C and inactive BP 1 include the orientation of the catalytic distal histidine, disruption of a hydrogen bond between this histidine and a conserved asparagine, and apparent substitution of calcium at the distal cation binding site with sodium at pH 7.5. These profound changes are a result of a dramatic structural rearrangement to the loop region between helices B and C. This is the first time that structural rearrangements linked to active site chemistry have been observed by crystallography in the peroxidase domain distal to heme.
- Department of Physical Chemistry, University of Copenhagen, København O, Denmark. anette@jerne.ki.ku.dk
Organizational Affiliation: