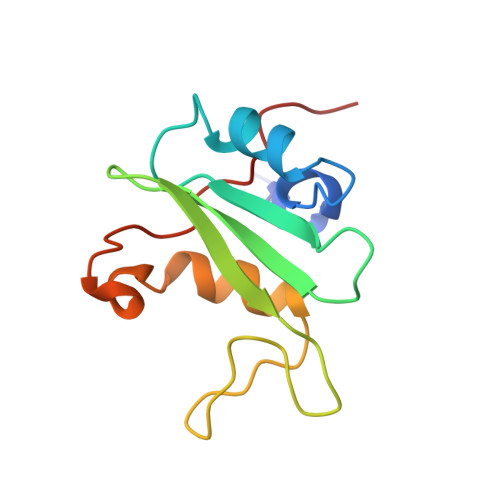
Solution structure of the C-terminal SH2 domain of the p85 alpha regulatory subunit of phosphoinositide 3-kinase.
Siegal, G., Davis, B., Kristensen, S.M., Sankar, A., Linacre, J., Stein, R.C., Panayotou, G., Waterfield, M.D., Driscoll, P.C.(1998) J Mol Biology 276: 461-478
- PubMed: 9512716 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1562
- Primary Citation Related Structures:
1BFI, 1BFJ - PubMed Abstract:
Heterodimeric class IA phosphoinositide 3-kinase (PI 3-kinase) plays a crucial role in a variety of cellular signalling events downstream of a number of cell-surface receptor tyrosine kinases. Activation of the enzyme is effected in part by the binding of two Src homology-2 domains (SH2) of the 85 kDa regulatory subunit to specific phosphotyrosine-containing peptide motifs within activated cytoplasmic receptor domains. The solution structure of the uncomplexed C-terminal SH2 (C-SH2) domain of the p85 alpha subunit of PI 3-kinase has been determined by means of multinuclear, double and triple-resonance NMR experiments and restrained molecular-dynamics simulated-annealing calculations. The solution structure clearly indicates that the uncomplexed C-SH2 domain conforms to the consensus polypeptide fold exhibited by other SH2 domains, with an additional short helical element at the N terminus. In particular, the C-SH2 structure is very similar to both the p85 alpha N-terminal SH2 domain (N-SH2) and the Src SH2 domain with a root mean square difference (rmsd) for 44 C alpha atoms of 1.09 and 0.89 A, respectively. The canonical BC, EF and BG loops are less well-defined by the experimental restraints and show greater variability in the ensemble of C-SH2 conformers. The lower level of definition in these regions may reflect the presence of conformational disorder, an interpretation supported by the absence or broadening of backbone and side-chain NMR resonances for some of these residues. NMR experiments were performed, where C-SH2 was titrated with phosphotyrosine-containing peptides corresponding to p85 alpha recognition sites in the cytoplasmic domain of the platelet-derived growth-factor receptor. The ligand-induced chemical-shift perturbations indicate the amino-acid residues in C-SH2 involved in peptide recognition follow the pattern predicted from homologous complexes. A series of C-SH2 mutants was generated and tested for phosphotyrosine peptide binding by surface plasmon resonance. Mutation of the invariant Arg36 (beta B5) to Met completely abolishes phosphopeptide binding. Mutation of each of Ser38, Ser39 or Lys40 in the BC loop to Ala reduces the affinity of C-SH2 for a cognate phosphopeptide, as does mutation of His93 (BG5) to Asn. These effects are consistent with the involvement of the BC loop and BG loops regions in ligation of phosphopeptide ligands. Mutation of Cys57 (beta D5) in C-SH2 to Ile, the corresponding residue type in the p85 alpha N-SH2 domain, results in a change in peptide binding selectivity of C-SH2 towards that demonstrated by p85 alpha N-SH2. This pattern of p85 alpha phosphopeptide binding specificity is interpreted in terms of a model of the p85 alpha/PDGF-receptor interaction.
- Ludwig Institute for Cancer Research, London, UK.
Organizational Affiliation: