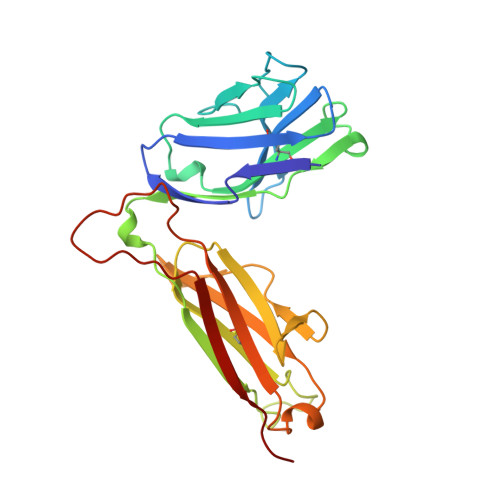
Crystal structure of the beta chain of a T cell antigen receptor.
Bentley, G.A., Boulot, G., Karjalainen, K., Mariuzza, R.A.(1995) Science 267: 1984-1987
- PubMed: 7701320 Search on PubMed
- DOI: https://doi.org/10.1126/science.7701320
- Primary Citation Related Structures:
1BEC - PubMed Abstract:
The crystal structure of the extracellular portion of the beta chain of a murine T cell antigen receptor (TCR), determined at a resolution of 1.7 angstroms, shows structural homology to immunoglobulins. The structure of the first and second hypervariable loops suggested that, in general, they adopt more restricted sets of conformations in TCR beta chains than those found in immunoglobulins; the third hypervariable loop had certain structural characteristics in common with those of immunoglobulin heavy chain variable domains. The variable and constant domains were in close contact, presumably restricting the flexibility of the beta chain. This may facilitate signal transduction from the TCR to the associated CD3 molecules in the TCR-CD3 complex.
- Unite d'Immunologie Structurale (CNRS URA 359), Institut Pasteur, Paris, France.
Organizational Affiliation: