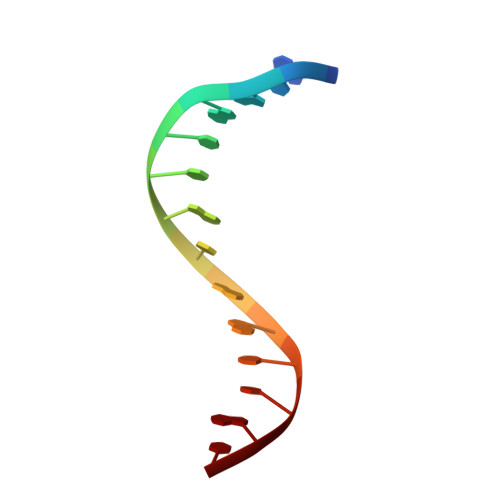
NMR structure of the extended Myb cognate sequence and modeling studies on specific DNA-Myb complexes.
Radha, P.K., Patel, P.K., Hosur, R.V.(1998) Biochemistry 37: 9952-9963
- PubMed: 9665700
- DOI: https://doi.org/10.1021/bi9806753
- Primary Citation of Related Structures:
1BDZ - PubMed Abstract:
The recognition sequence of the Myb protein has been recently described to be pyAACKGHH (where py = T/C, K = G/T, and H = A/C/T), modifying the earlier identification as pyAACKG [Ording, E., et al. (1994) Eur. J. Biochem. 222, 113-120]. We had earlier determined the solution structure of the minimal cognate sequence TAACGG, choosing py = T and K = G, embeded in a 12-mer DNA duplex by NMR and related computational techniques [Radha, P. K., et al. (1995) Biochemistry 34, 5913-5912]. To understand the structural significance of the above modification and the role of the variability in the recognition sequence, we have investigated here the solution structure of a different DNA segment, d-ACAACTGCAGTTGT, which contains the extended Myb cognate site, CAACTGCA. The three-dimensional structure of the 14-mer duplex has been determined from NMR data by relaxation matrix and restrained molecular dynamics calculations. The structure of the above cognate sequence in the 14-mer duplex has been compared with that of the cognate sequence, TAACGG, in the 12-mer duplex and also with that in the NMR structure of the Myb DNA binding domain (R2R3)-DNA complex determined by Ogata et al. recently [Ogata, K., et al. (1994) Cell 79, 639-648]. The comparison highlighted differences in several structural parameters for the cognate sites in the DNA segments. Modeling studies by taking out the protein from the complex and presenting it with 12-mer and 14-mer DNA structures indicated that the protein induces structural alterations to drive the cognate site to a reasonably conserved structure. The extent of similarity of the derived structures was, however, dependent on the base sequences. Base changes in the minimal cognate sequence in the 12-mer-protein complex and in the 14-mer-protein complex so as to match the sequence of Ogata et al. produced a more conserved structure of the complex. A reverse exercise, in which the Ogata DNA in the complex was mutated to match the 12-mer and 14-mer minimal cognate sequences, complemented the above observations of the subtle sequence dependence of the structure in the complex. On the other hand, base changes in the extension did not influence the DNA-protein complex structure significantly. We also observed that the structural changes in the protein were very minor when different DNA sequences or different DNA structures were presented to it. These observations would be of interest from the point of view of DNA-Myb recognition.
- Department of Chemical Sciences, Tata Institute of Fundamental Research, Mumbai, India.
Organizational Affiliation: