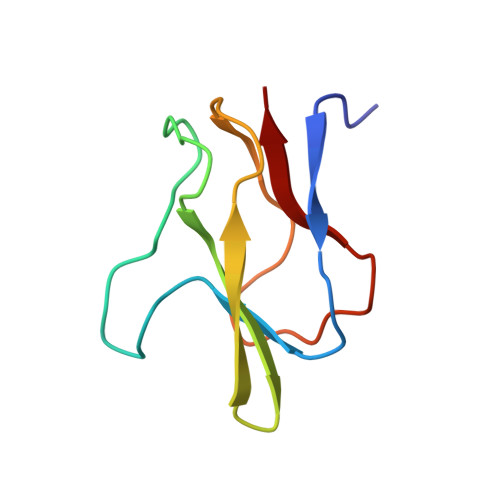
Structure of the biotinyl domain of acetyl-coenzyme A carboxylase determined by MAD phasing.
Athappilly, F.K., Hendrickson, W.A.(1995) Structure 3: 1407-1419
- PubMed: 8747466
- DOI: https://doi.org/10.1016/s0969-2126(01)00277-5
- Primary Citation Related Structures:
1BDO - PubMed Abstract:
Acetyl-coenzyme A carboxylase catalyzes the first committed step of fatty acid biosynthesis. Universally, this reaction involves three functional components all related to a carboxybiotinyl intermediate. A biotinyl domain shuttles its covalently attached biotin prosthetic group between the active sites of a biotin carboxylase and a carboxyl transferase. In Escherichia coli, the three components reside in separate subunits: a biotinyl domain is the functional portion of one of these, biotin carboxy carrier protein (BCCP). We have expressed natural and selenomethionyl (Se-met) BCCP from E. coli as biotinylated recombinant proteins, proteolyzed them with subtilisin Carlsberg to produce the biotinyl domains BCCP and Se-met BCCPsc, determined the crystal structure of Se-met BCCPsc using a modified version of the multiwavelength anomalous diffraction (MAD) phasing protocol, and refined the structure for the natural BCCPsc at 1.8 A resolution. The structure may be described as a capped beta sandwich with quasi-dyad symmetry. Each half contains a characteristic hammerhead motif. The biotinylated lysin is located at a hairpin beta turn which connects the two symmetric halves of the molecule, and its biotinyl group interacts with a non-symmetric protrusion from the core. This first crystal structure of a biotinyl domain helps to unravel the central role of such domains in reactions catalyzed by biotin-dependent carboxylases. The hammerhead structure observed twice in BCCPsc may be regarded as the basic structural motif of biotinyl and lipoyl domains of a superfamily of enzymes. The new MAD phasing techniques developed in the course of determining this structure enhance the power of the MAD method.
- Department of Biochemistry and Molecular Biophysics, College of Physicians and Surgeons of Columbia University, New York, NY 10032, USA.
Organizational Affiliation: