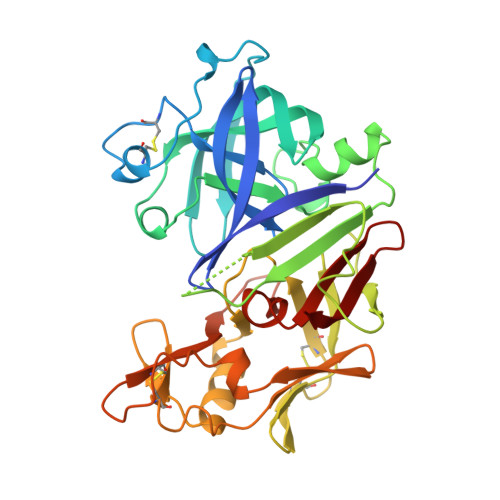
X-ray analyses of peptide-inhibitor complexes define the structural basis of specificity for human and mouse renins.
Dhanaraj, V., Dealwis, C.G., Frazao, C., Badasso, M., Sibanda, B.L., Tickle, I.J., Cooper, J.B., Driessen, H.P., Newman, M., Aguilar, C., Wood, S.P., Blundell, T.L., Hobart, P.M., Geoghegan, K.F., Ammirati, M.J., Danley, D.E.(1992) Nature 357: 466-472
- PubMed: 1608447 Search on PubMed
- DOI: https://doi.org/10.1038/357466a0
- Primary Citation Related Structures:
1BBS - PubMed Abstract:
X-ray analyses have defined the three-dimensional structures of crystals of mouse and human renins complexed with peptide inhibitors at resolutions of 1.9 and 2.8 A, respectively. The exquisite specificity of renin arises partly from ordered loop regions at the periphery of the binding cleft. Although the pattern of main-chain hydrogen bonding in other aspartic proteinase inhibitor complexes is conserved in renins, differences in the positions of secondary structure elements (particularly helices) also lead to improved specificity in renins for angiotensinogen substrates.
- Department of Crystallography, Birkbeck College, London, UK.
Organizational Affiliation: