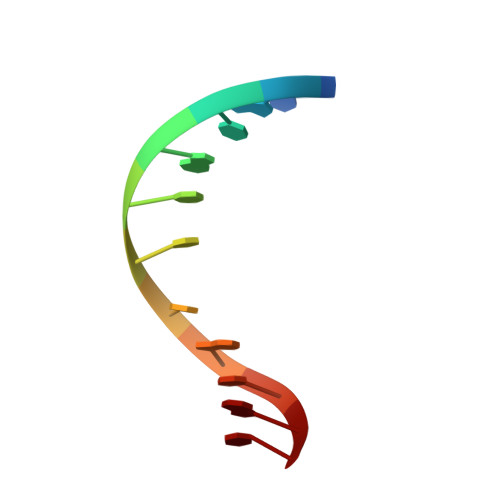
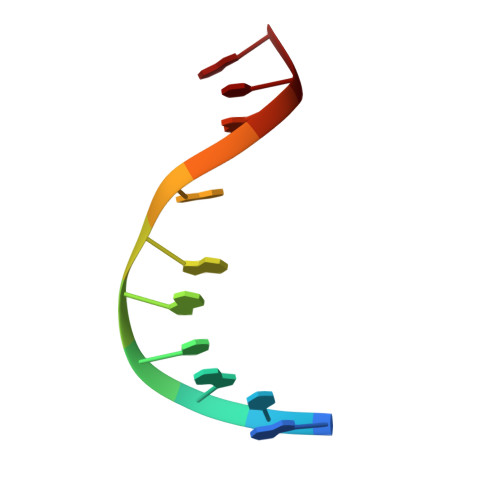
Solution structure of an oligodeoxynucleotide duplex containing the exocyclic lesion 3,N4-etheno-2'-deoxycytidine opposite 2'-deoxyadenosine, determined by NMR spectroscopy and restrained molecular dynamics.
Korobka, A., Cullinan, D., Cosman, M., Grollman, A.P., Patel, D.J., Eisenberg, M., de los Santos, C.(1996) Biochemistry 35: 13310-13318
- PubMed: 8873597 Search on PubMed
- DOI: https://doi.org/10.1021/bi9605696
- Primary Citation Related Structures:
1B6Y - PubMed Abstract:
The d(C-G-T-A-C-epsilon C-C-A-T-G-C).d(G-C-A-T-G-A-G-T-A-C-G) oligodeoxynucleotide duplex containing the 3, N4-etheno-2'-deoxycytidine adduct positioned opposite 2'-deoxyadenosine in the center of the helix has been analyzed by proton NMR spectroscopy and restrained molecular dynamics. The spectroscopic data establish a right-handed duplex, with sugar puckers in the C2'-endo/C3'-exo range, residues adopting an anti conformation around the glycosidic torsion angle and, with the exception of epsilon C.dA, Watson-Crick hydrogen bond alignment for all base pairs. Molecular dynamics simulations, restrained by the full relaxation matrix approach, produced a three-dimensional model with an NMR R-factor of 7%. The duplex structure shows no significant perturbation of the sugar-phosphate backbone, which remains in B-form. The exocyclic adduct and its partner dA are incorporated into the helix without producing a noticeable kink. The epsilon C.dA alignment adopts a staggered conformation with each residue displaced toward the 5'-terminus and intercalated between bases on the opposite strand, without increase of inter-phosphate distances. The partial intercalation of the epsilon C (anti).dA(anti) alignment allows stacking between the aromatic rings of epsilon C and dA and with base pairs adjacent to the lesion, suggesting an important role played by hydrophobic forces in the stabilization of the solution structure.
- Department of Pharmacological Sciences, State University of New York at Stony Brook 11794-8651, USA.
Organizational Affiliation: