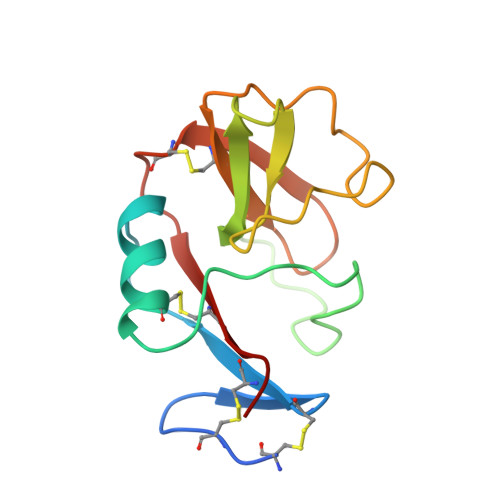
Structure of CD94 reveals a novel C-type lectin fold: implications for the NK cell-associated CD94/NKG2 receptors.
Boyington, J.C., Riaz, A.N., Patamawenu, A., Coligan, J.E., Brooks, A.G., Sun, P.D.(1999) Immunity 10: 75-82
- PubMed: 10023772
- DOI: https://doi.org/10.1016/s1074-7613(00)80008-4
- Primary Citation of Related Structures:
1B6E - PubMed Abstract:
The crystal structure of the extracellular domain of CD94, a component of the CD94/NKG2 NK cell receptor, has been determined to 2.6 A resolution, revealing a unique variation of the C-type lectin fold. In this variation, the second alpha helix, corresponding to residues 102-112, is replaced by a loop, the putative carbohydrate-binding site is significantly altered, and the Ca2+-binding site appears nonfunctional. This structure may serve as a prototype for other NK cell receptors such as Ly-49, NKR-P1, and CD69. The CD94 dimer observed in the crystal has an extensive hydrophobic interface that stabilizes the loop conformation of residues 102-112. The formation of this dimer reveals a putative ligand-binding region for HLA-E and suggests how NKG2 interacts with CD94.
- Structural Biology Section, Office of the Scientific Director, National Institute of Allergy and Infectious Diseases, Rockville, Maryland 20852, USA.
Organizational Affiliation: