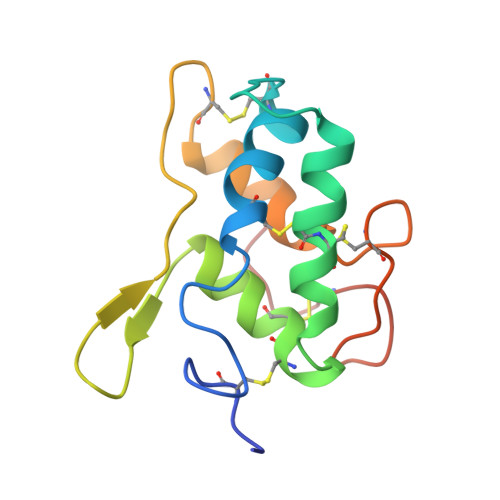
Structure of the bifunctional inhibitor of trypsin and alpha-amylase from ragi seeds at 2.2 A resolution.
Gourinath, S., Alam, N., Srinivasan, A., Betzel, C., Singh, T.P.(2000) Acta Crystallogr D Biol Crystallogr 56: 287-293
- PubMed: 10713515 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444999016601
- Primary Citation Related Structures:
1B1U - PubMed Abstract:
The crystal structure of a bifunctional inhibitor of alpha-amylase and trypsin (RATI) from ragi seeds (Indian finger millet, Eleusine coracana Gaertneri) has been determined by X-ray diffraction at 2.2 A resolution. The inhibitor consists of 122 amino acids, with five disulfide bridges, and belongs to the plant alpha-amylase/trypsin inhibitor family. The crystals were grown by the microdialysis method using ammonium sulfate as a precipitating agent. The structure was determined by the molecular-replacement method using as models the structures of Corn Hageman factor inhibitor (CHFI) and of RATI at 2.9 A resolution determined previously. It has been refined to an R factor of 21.9%. The structure shows an r.m.s. deviation for C(alpha) atoms of 2.0 A compared with its own NMR structure, whereas the corresponding value compared with CHFI is found to be 1.4 A. The r.m.s. difference for C(alpha) atoms when compared with the same protein in the structure of the complex with alpha-amylase is 0.7 A. The conformations of trypsin-binding loop and the alpha-amylase-binding N-terminal region were also found to be similar in the crystal structures of native RATI and its complex with alpha-amylase. These regions differed considerably in the NMR structure.
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi-110029, India.
Organizational Affiliation: